### What changes were proposed in this pull request?
This PR proposes to add one more newline to clearly separate JVM and Python tracebacks:
Before:
```
Traceback (most recent call last):
...
pyspark.sql.utils.AnalysisException: Reference 'column' is ambiguous, could be: column, column.;
JVM stacktrace:
org.apache.spark.sql.AnalysisException: Reference 'column' is ambiguous, could be: column, column.;
...
```
After:
```
Traceback (most recent call last):
...
pyspark.sql.utils.AnalysisException: Reference 'column' is ambiguous, could be: column, column.;
JVM stacktrace:
org.apache.spark.sql.AnalysisException: Reference 'column' is ambiguous, could be: column, column.;
...
```
This is kind of a followup of e69466056f (SPARK-31849).
### Why are the changes needed?
To make it easier to read.
### Does this PR introduce _any_ user-facing change?
It's in the unreleased branches.
### How was this patch tested?
Manually tested.
Closes#28732 from HyukjinKwon/python-minor.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
Scala:
```scala
scala> spark.range(10).explain("cost")
```
```
== Optimized Logical Plan ==
Range (0, 10, step=1, splits=Some(12)), Statistics(sizeInBytes=80.0 B)
== Physical Plan ==
*(1) Range (0, 10, step=1, splits=12)
```
PySpark:
```python
>>> spark.range(10).explain("cost")
```
```
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "/.../spark/python/pyspark/sql/dataframe.py", line 333, in explain
raise TypeError(err_msg)
TypeError: extended (optional) should be provided as bool, got <class 'str'>
```
In addition, it is consistent with other codes too, for example, `DataFrame.sample` also can support `DataFrame.sample(1.0)` and `DataFrame.sample(False)`.
### Why are the changes needed?
To provide the consistent API support across APIs.
### Does this PR introduce _any_ user-facing change?
Nope, it's only changes in unreleased branches.
If this lands to master only, yes, users will be able to set `mode` as `df.explain("...")` in Spark 3.1.
After this PR:
```python
>>> spark.range(10).explain("cost")
```
```
== Optimized Logical Plan ==
Range (0, 10, step=1, splits=Some(12)), Statistics(sizeInBytes=80.0 B)
== Physical Plan ==
*(1) Range (0, 10, step=1, splits=12)
```
### How was this patch tested?
Unittest was added and manually tested as well to make sure:
```python
spark.range(10).explain(True)
spark.range(10).explain(False)
spark.range(10).explain("cost")
spark.range(10).explain(extended="cost")
spark.range(10).explain(mode="cost")
spark.range(10).explain()
spark.range(10).explain(True, "cost")
spark.range(10).explain(1.0)
```
Closes#28711 from HyukjinKwon/SPARK-31895.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
It increases the timeout for `StreamingLinearRegressionWithTests.test_train_prediction`
```
Traceback (most recent call last):
File "/home/jenkins/workspace/SparkPullRequestBuilder3/python/pyspark/mllib/tests/test_streaming_algorithms.py", line 503, in test_train_prediction
self._eventually(condition)
File "/home/jenkins/workspace/SparkPullRequestBuilder3/python/pyspark/mllib/tests/test_streaming_algorithms.py", line 69, in _eventually
lastValue = condition()
File "/home/jenkins/workspace/SparkPullRequestBuilder3/python/pyspark/mllib/tests/test_streaming_algorithms.py", line 498, in condition
self.assertGreater(errors[1] - errors[-1], 2)
AssertionError: 1.672640157855923 not greater than 2
```
This could likely happen when the PySpark tests run in parallel and it become slow.
### Why are the changes needed?
To make the tests less flaky. Seems it's being reported multiple times:
https://amplab.cs.berkeley.edu/jenkins/job/SparkPullRequestBuilder/123144/consoleFullhttps://amplab.cs.berkeley.edu/jenkins/job/SparkPullRequestBuilder/123146/testReport/https://amplab.cs.berkeley.edu/jenkins/job/SparkPullRequestBuilder/123141/testReport/https://amplab.cs.berkeley.edu/jenkins/job/SparkPullRequestBuilder/123142/testReport/
### Does this PR introduce _any_ user-facing change?
No, test-only.
### How was this patch tested?
Jenkins will test it out.
Closes#28701 from HyukjinKwon/SPARK-29137.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: Dongjoon Hyun <dongjoon@apache.org>
### What changes were proposed in this pull request?
This PR proposes to make PySpark exception more Pythonic by hiding JVM stacktrace by default. It can be enabled by turning on `spark.sql.pyspark.jvmStacktrace.enabled` configuration.
```
Traceback (most recent call last):
...
pyspark.sql.utils.PythonException:
An exception was thrown from Python worker in the executor. The below is the Python worker stacktrace.
Traceback (most recent call last):
...
```
If this `spark.sql.pyspark.jvmStacktrace.enabled` is enabled, it appends:
```
JVM stacktrace:
org.apache.spark.Exception: ...
...
```
For example, the codes below:
```python
from pyspark.sql.functions import udf
udf
def divide_by_zero(v):
raise v / 0
spark.range(1).select(divide_by_zero("id")).show()
```
will show an error messages that looks like Python exception thrown from the local.
<details>
<summary>Python exception message when <code>spark.sql.pyspark.jvmStacktrace.enabled</code> is off (default)</summary>
```
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "/.../spark/python/pyspark/sql/dataframe.py", line 427, in show
print(self._jdf.showString(n, 20, vertical))
File "/.../spark/python/lib/py4j-0.10.9-src.zip/py4j/java_gateway.py", line 1305, in __call__
File "/.../spark/python/pyspark/sql/utils.py", line 131, in deco
raise_from(converted)
File "<string>", line 3, in raise_from
pyspark.sql.utils.PythonException:
An exception was thrown from Python worker in the executor. The below is the Python worker stacktrace.
Traceback (most recent call last):
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 605, in main
process()
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 597, in process
serializer.dump_stream(out_iter, outfile)
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 223, in dump_stream
self.serializer.dump_stream(self._batched(iterator), stream)
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 141, in dump_stream
for obj in iterator:
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 212, in _batched
for item in iterator:
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 450, in mapper
result = tuple(f(*[a[o] for o in arg_offsets]) for (arg_offsets, f) in udfs)
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 450, in <genexpr>
result = tuple(f(*[a[o] for o in arg_offsets]) for (arg_offsets, f) in udfs)
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 90, in <lambda>
return lambda *a: f(*a)
File "/.../spark/python/lib/pyspark.zip/pyspark/util.py", line 107, in wrapper
return f(*args, **kwargs)
File "<stdin>", line 3, in divide_by_zero
ZeroDivisionError: division by zero
```
</details>
<details>
<summary>Python exception message when <code>spark.sql.pyspark.jvmStacktrace.enabled</code> is on</summary>
```
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "/.../spark/python/pyspark/sql/dataframe.py", line 427, in show
print(self._jdf.showString(n, 20, vertical))
File "/.../spark/python/lib/py4j-0.10.9-src.zip/py4j/java_gateway.py", line 1305, in __call__
File "/.../spark/python/pyspark/sql/utils.py", line 137, in deco
raise_from(converted)
File "<string>", line 3, in raise_from
pyspark.sql.utils.PythonException:
An exception was thrown from Python worker in the executor. The below is the Python worker stacktrace.
Traceback (most recent call last):
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 605, in main
process()
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 597, in process
serializer.dump_stream(out_iter, outfile)
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 223, in dump_stream
self.serializer.dump_stream(self._batched(iterator), stream)
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 141, in dump_stream
for obj in iterator:
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 212, in _batched
for item in iterator:
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 450, in mapper
result = tuple(f(*[a[o] for o in arg_offsets]) for (arg_offsets, f) in udfs)
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 450, in <genexpr>
result = tuple(f(*[a[o] for o in arg_offsets]) for (arg_offsets, f) in udfs)
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 90, in <lambda>
return lambda *a: f(*a)
File "/.../spark/python/lib/pyspark.zip/pyspark/util.py", line 107, in wrapper
return f(*args, **kwargs)
File "<stdin>", line 3, in divide_by_zero
ZeroDivisionError: division by zero
JVM stacktrace:
org.apache.spark.SparkException: Job aborted due to stage failure: Task 0 in stage 1.0 failed 4 times, most recent failure: Lost task 0.3 in stage 1.0 (TID 4, 192.168.35.193, executor 0): org.apache.spark.api.python.PythonException: Traceback (most recent call last):
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 605, in main
process()
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 597, in process
serializer.dump_stream(out_iter, outfile)
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 223, in dump_stream
self.serializer.dump_stream(self._batched(iterator), stream)
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 141, in dump_stream
for obj in iterator:
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 212, in _batched
for item in iterator:
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 450, in mapper
result = tuple(f(*[a[o] for o in arg_offsets]) for (arg_offsets, f) in udfs)
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 450, in <genexpr>
result = tuple(f(*[a[o] for o in arg_offsets]) for (arg_offsets, f) in udfs)
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 90, in <lambda>
return lambda *a: f(*a)
File "/.../spark/python/lib/pyspark.zip/pyspark/util.py", line 107, in wrapper
return f(*args, **kwargs)
File "<stdin>", line 3, in divide_by_zero
ZeroDivisionError: division by zero
at org.apache.spark.api.python.BasePythonRunner$ReaderIterator.handlePythonException(PythonRunner.scala:516)
at org.apache.spark.sql.execution.python.PythonUDFRunner$$anon$2.read(PythonUDFRunner.scala:81)
at org.apache.spark.sql.execution.python.PythonUDFRunner$$anon$2.read(PythonUDFRunner.scala:64)
at org.apache.spark.api.python.BasePythonRunner$ReaderIterator.hasNext(PythonRunner.scala:469)
at org.apache.spark.InterruptibleIterator.hasNext(InterruptibleIterator.scala:37)
at scala.collection.Iterator$$anon$11.hasNext(Iterator.scala:489)
at scala.collection.Iterator$$anon$10.hasNext(Iterator.scala:458)
at scala.collection.Iterator$$anon$10.hasNext(Iterator.scala:458)
at org.apache.spark.sql.catalyst.expressions.GeneratedClass$GeneratedIteratorForCodegenStage2.processNext(Unknown Source)
at org.apache.spark.sql.execution.BufferedRowIterator.hasNext(BufferedRowIterator.java:43)
at org.apache.spark.sql.execution.WholeStageCodegenExec$$anon$1.hasNext(WholeStageCodegenExec.scala:753)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$getByteArrayRdd$1(SparkPlan.scala:340)
at org.apache.spark.rdd.RDD.$anonfun$mapPartitionsInternal$2(RDD.scala:898)
at org.apache.spark.rdd.RDD.$anonfun$mapPartitionsInternal$2$adapted(RDD.scala:898)
at org.apache.spark.rdd.MapPartitionsRDD.compute(MapPartitionsRDD.scala:52)
at org.apache.spark.rdd.RDD.computeOrReadCheckpoint(RDD.scala:373)
at org.apache.spark.rdd.RDD.iterator(RDD.scala:337)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:127)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:469)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1377)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:472)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
at java.lang.Thread.run(Thread.java:748)
Driver stacktrace:
at org.apache.spark.scheduler.DAGScheduler.failJobAndIndependentStages(DAGScheduler.scala:2117)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2(DAGScheduler.scala:2066)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2$adapted(DAGScheduler.scala:2065)
at scala.collection.mutable.ResizableArray.foreach(ResizableArray.scala:62)
at scala.collection.mutable.ResizableArray.foreach$(ResizableArray.scala:55)
at scala.collection.mutable.ArrayBuffer.foreach(ArrayBuffer.scala:49)
at org.apache.spark.scheduler.DAGScheduler.abortStage(DAGScheduler.scala:2065)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1(DAGScheduler.scala:1021)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1$adapted(DAGScheduler.scala:1021)
at scala.Option.foreach(Option.scala:407)
at org.apache.spark.scheduler.DAGScheduler.handleTaskSetFailed(DAGScheduler.scala:1021)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.doOnReceive(DAGScheduler.scala:2297)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2246)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2235)
at org.apache.spark.util.EventLoop$$anon$1.run(EventLoop.scala:49)
at org.apache.spark.scheduler.DAGScheduler.runJob(DAGScheduler.scala:823)
at org.apache.spark.SparkContext.runJob(SparkContext.scala:2108)
at org.apache.spark.SparkContext.runJob(SparkContext.scala:2129)
at org.apache.spark.SparkContext.runJob(SparkContext.scala:2148)
at org.apache.spark.sql.execution.SparkPlan.executeTake(SparkPlan.scala:467)
at org.apache.spark.sql.execution.SparkPlan.executeTake(SparkPlan.scala:420)
at org.apache.spark.sql.execution.CollectLimitExec.executeCollect(limit.scala:47)
at org.apache.spark.sql.Dataset.collectFromPlan(Dataset.scala:3653)
at org.apache.spark.sql.Dataset.$anonfun$head$1(Dataset.scala:2695)
at org.apache.spark.sql.Dataset.$anonfun$withAction$1(Dataset.scala:3644)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$5(SQLExecution.scala:103)
at org.apache.spark.sql.execution.SQLExecution$.withSQLConfPropagated(SQLExecution.scala:163)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$1(SQLExecution.scala:90)
at org.apache.spark.sql.SparkSession.withActive(SparkSession.scala:763)
at org.apache.spark.sql.execution.SQLExecution$.withNewExecutionId(SQLExecution.scala:64)
at org.apache.spark.sql.Dataset.withAction(Dataset.scala:3642)
at org.apache.spark.sql.Dataset.head(Dataset.scala:2695)
at org.apache.spark.sql.Dataset.take(Dataset.scala:2902)
at org.apache.spark.sql.Dataset.getRows(Dataset.scala:300)
at org.apache.spark.sql.Dataset.showString(Dataset.scala:337)
at sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
at sun.reflect.NativeMethodAccessorImpl.invoke(NativeMethodAccessorImpl.java:62)
at sun.reflect.DelegatingMethodAccessorImpl.invoke(DelegatingMethodAccessorImpl.java:43)
at java.lang.reflect.Method.invoke(Method.java:498)
at py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
at py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:357)
at py4j.Gateway.invoke(Gateway.java:282)
at py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
at py4j.commands.CallCommand.execute(CallCommand.java:79)
at py4j.GatewayConnection.run(GatewayConnection.java:238)
at java.lang.Thread.run(Thread.java:748)
Caused by: org.apache.spark.api.python.PythonException: Traceback (most recent call last):
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 605, in main
process()
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 597, in process
serializer.dump_stream(out_iter, outfile)
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 223, in dump_stream
self.serializer.dump_stream(self._batched(iterator), stream)
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 141, in dump_stream
for obj in iterator:
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 212, in _batched
for item in iterator:
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 450, in mapper
result = tuple(f(*[a[o] for o in arg_offsets]) for (arg_offsets, f) in udfs)
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 450, in <genexpr>
result = tuple(f(*[a[o] for o in arg_offsets]) for (arg_offsets, f) in udfs)
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 90, in <lambda>
return lambda *a: f(*a)
File "/.../spark/python/lib/pyspark.zip/pyspark/util.py", line 107, in wrapper
return f(*args, **kwargs)
File "<stdin>", line 3, in divide_by_zero
ZeroDivisionError: division by zero
at org.apache.spark.api.python.BasePythonRunner$ReaderIterator.handlePythonException(PythonRunner.scala:516)
at org.apache.spark.sql.execution.python.PythonUDFRunner$$anon$2.read(PythonUDFRunner.scala:81)
at org.apache.spark.sql.execution.python.PythonUDFRunner$$anon$2.read(PythonUDFRunner.scala:64)
at org.apache.spark.api.python.BasePythonRunner$ReaderIterator.hasNext(PythonRunner.scala:469)
at org.apache.spark.InterruptibleIterator.hasNext(InterruptibleIterator.scala:37)
at scala.collection.Iterator$$anon$11.hasNext(Iterator.scala:489)
at scala.collection.Iterator$$anon$10.hasNext(Iterator.scala:458)
at scala.collection.Iterator$$anon$10.hasNext(Iterator.scala:458)
at org.apache.spark.sql.catalyst.expressions.GeneratedClass$GeneratedIteratorForCodegenStage2.processNext(Unknown Source)
at org.apache.spark.sql.execution.BufferedRowIterator.hasNext(BufferedRowIterator.java:43)
at org.apache.spark.sql.execution.WholeStageCodegenExec$$anon$1.hasNext(WholeStageCodegenExec.scala:753)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$getByteArrayRdd$1(SparkPlan.scala:340)
at org.apache.spark.rdd.RDD.$anonfun$mapPartitionsInternal$2(RDD.scala:898)
at org.apache.spark.rdd.RDD.$anonfun$mapPartitionsInternal$2$adapted(RDD.scala:898)
at org.apache.spark.rdd.MapPartitionsRDD.compute(MapPartitionsRDD.scala:52)
at org.apache.spark.rdd.RDD.computeOrReadCheckpoint(RDD.scala:373)
at org.apache.spark.rdd.RDD.iterator(RDD.scala:337)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:127)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:469)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1377)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:472)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
... 1 more
```
</details>
<details>
<summary>Python exception message without this change</summary>
```
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "/.../spark/python/pyspark/sql/dataframe.py", line 427, in show
print(self._jdf.showString(n, 20, vertical))
File "/.../spark/python/lib/py4j-0.10.9-src.zip/py4j/java_gateway.py", line 1305, in __call__
File "/.../spark/python/pyspark/sql/utils.py", line 98, in deco
return f(*a, **kw)
File "/.../spark/python/lib/py4j-0.10.9-src.zip/py4j/protocol.py", line 328, in get_return_value
py4j.protocol.Py4JJavaError: An error occurred while calling o160.showString.
: org.apache.spark.SparkException: Job aborted due to stage failure: Task 10 in stage 5.0 failed 4 times, most recent failure: Lost task 10.3 in stage 5.0 (TID 37, 192.168.35.193, executor 3): org.apache.spark.api.python.PythonException: Traceback (most recent call last):
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 605, in main
process()
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 597, in process
serializer.dump_stream(out_iter, outfile)
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 223, in dump_stream
self.serializer.dump_stream(self._batched(iterator), stream)
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 141, in dump_stream
for obj in iterator:
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 212, in _batched
for item in iterator:
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 450, in mapper
result = tuple(f(*[a[o] for o in arg_offsets]) for (arg_offsets, f) in udfs)
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 450, in <genexpr>
result = tuple(f(*[a[o] for o in arg_offsets]) for (arg_offsets, f) in udfs)
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 90, in <lambda>
return lambda *a: f(*a)
File "/.../spark/python/lib/pyspark.zip/pyspark/util.py", line 107, in wrapper
return f(*args, **kwargs)
File "<stdin>", line 3, in divide_by_zero
ZeroDivisionError: division by zero
at org.apache.spark.api.python.BasePythonRunner$ReaderIterator.handlePythonException(PythonRunner.scala:516)
at org.apache.spark.sql.execution.python.PythonUDFRunner$$anon$2.read(PythonUDFRunner.scala:81)
at org.apache.spark.sql.execution.python.PythonUDFRunner$$anon$2.read(PythonUDFRunner.scala:64)
at org.apache.spark.api.python.BasePythonRunner$ReaderIterator.hasNext(PythonRunner.scala:469)
at org.apache.spark.InterruptibleIterator.hasNext(InterruptibleIterator.scala:37)
at scala.collection.Iterator$$anon$11.hasNext(Iterator.scala:489)
at scala.collection.Iterator$$anon$10.hasNext(Iterator.scala:458)
at scala.collection.Iterator$$anon$10.hasNext(Iterator.scala:458)
at org.apache.spark.sql.catalyst.expressions.GeneratedClass$GeneratedIteratorForCodegenStage2.processNext(Unknown Source)
at org.apache.spark.sql.execution.BufferedRowIterator.hasNext(BufferedRowIterator.java:43)
at org.apache.spark.sql.execution.WholeStageCodegenExec$$anon$1.hasNext(WholeStageCodegenExec.scala:753)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$getByteArrayRdd$1(SparkPlan.scala:340)
at org.apache.spark.rdd.RDD.$anonfun$mapPartitionsInternal$2(RDD.scala:898)
at org.apache.spark.rdd.RDD.$anonfun$mapPartitionsInternal$2$adapted(RDD.scala:898)
at org.apache.spark.rdd.MapPartitionsRDD.compute(MapPartitionsRDD.scala:52)
at org.apache.spark.rdd.RDD.computeOrReadCheckpoint(RDD.scala:373)
at org.apache.spark.rdd.RDD.iterator(RDD.scala:337)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:127)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:469)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1377)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:472)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
at java.lang.Thread.run(Thread.java:748)
Driver stacktrace:
at org.apache.spark.scheduler.DAGScheduler.failJobAndIndependentStages(DAGScheduler.scala:2117)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2(DAGScheduler.scala:2066)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2$adapted(DAGScheduler.scala:2065)
at scala.collection.mutable.ResizableArray.foreach(ResizableArray.scala:62)
at scala.collection.mutable.ResizableArray.foreach$(ResizableArray.scala:55)
at scala.collection.mutable.ArrayBuffer.foreach(ArrayBuffer.scala:49)
at org.apache.spark.scheduler.DAGScheduler.abortStage(DAGScheduler.scala:2065)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1(DAGScheduler.scala:1021)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1$adapted(DAGScheduler.scala:1021)
at scala.Option.foreach(Option.scala:407)
at org.apache.spark.scheduler.DAGScheduler.handleTaskSetFailed(DAGScheduler.scala:1021)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.doOnReceive(DAGScheduler.scala:2297)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2246)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2235)
at org.apache.spark.util.EventLoop$$anon$1.run(EventLoop.scala:49)
at org.apache.spark.scheduler.DAGScheduler.runJob(DAGScheduler.scala:823)
at org.apache.spark.SparkContext.runJob(SparkContext.scala:2108)
at org.apache.spark.SparkContext.runJob(SparkContext.scala:2129)
at org.apache.spark.SparkContext.runJob(SparkContext.scala:2148)
at org.apache.spark.sql.execution.SparkPlan.executeTake(SparkPlan.scala:467)
at org.apache.spark.sql.execution.SparkPlan.executeTake(SparkPlan.scala:420)
at org.apache.spark.sql.execution.CollectLimitExec.executeCollect(limit.scala:47)
at org.apache.spark.sql.Dataset.collectFromPlan(Dataset.scala:3653)
at org.apache.spark.sql.Dataset.$anonfun$head$1(Dataset.scala:2695)
at org.apache.spark.sql.Dataset.$anonfun$withAction$1(Dataset.scala:3644)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$5(SQLExecution.scala:103)
at org.apache.spark.sql.execution.SQLExecution$.withSQLConfPropagated(SQLExecution.scala:163)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$1(SQLExecution.scala:90)
at org.apache.spark.sql.SparkSession.withActive(SparkSession.scala:763)
at org.apache.spark.sql.execution.SQLExecution$.withNewExecutionId(SQLExecution.scala:64)
at org.apache.spark.sql.Dataset.withAction(Dataset.scala:3642)
at org.apache.spark.sql.Dataset.head(Dataset.scala:2695)
at org.apache.spark.sql.Dataset.take(Dataset.scala:2902)
at org.apache.spark.sql.Dataset.getRows(Dataset.scala:300)
at org.apache.spark.sql.Dataset.showString(Dataset.scala:337)
at sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
at sun.reflect.NativeMethodAccessorImpl.invoke(NativeMethodAccessorImpl.java:62)
at sun.reflect.DelegatingMethodAccessorImpl.invoke(DelegatingMethodAccessorImpl.java:43)
at java.lang.reflect.Method.invoke(Method.java:498)
at py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
at py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:357)
at py4j.Gateway.invoke(Gateway.java:282)
at py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
at py4j.commands.CallCommand.execute(CallCommand.java:79)
at py4j.GatewayConnection.run(GatewayConnection.java:238)
at java.lang.Thread.run(Thread.java:748)
Caused by: org.apache.spark.api.python.PythonException: Traceback (most recent call last):
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 605, in main
process()
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 597, in process
serializer.dump_stream(out_iter, outfile)
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 223, in dump_stream
self.serializer.dump_stream(self._batched(iterator), stream)
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 141, in dump_stream
for obj in iterator:
File "/.../spark/python/lib/pyspark.zip/pyspark/serializers.py", line 212, in _batched
for item in iterator:
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 450, in mapper
result = tuple(f(*[a[o] for o in arg_offsets]) for (arg_offsets, f) in udfs)
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 450, in <genexpr>
result = tuple(f(*[a[o] for o in arg_offsets]) for (arg_offsets, f) in udfs)
File "/.../spark/python/lib/pyspark.zip/pyspark/worker.py", line 90, in <lambda>
return lambda *a: f(*a)
File "/.../spark/python/lib/pyspark.zip/pyspark/util.py", line 107, in wrapper
return f(*args, **kwargs)
File "<stdin>", line 3, in divide_by_zero
ZeroDivisionError: division by zero
at org.apache.spark.api.python.BasePythonRunner$ReaderIterator.handlePythonException(PythonRunner.scala:516)
at org.apache.spark.sql.execution.python.PythonUDFRunner$$anon$2.read(PythonUDFRunner.scala:81)
at org.apache.spark.sql.execution.python.PythonUDFRunner$$anon$2.read(PythonUDFRunner.scala:64)
at org.apache.spark.api.python.BasePythonRunner$ReaderIterator.hasNext(PythonRunner.scala:469)
at org.apache.spark.InterruptibleIterator.hasNext(InterruptibleIterator.scala:37)
at scala.collection.Iterator$$anon$11.hasNext(Iterator.scala:489)
at scala.collection.Iterator$$anon$10.hasNext(Iterator.scala:458)
at scala.collection.Iterator$$anon$10.hasNext(Iterator.scala:458)
at org.apache.spark.sql.catalyst.expressions.GeneratedClass$GeneratedIteratorForCodegenStage2.processNext(Unknown Source)
at org.apache.spark.sql.execution.BufferedRowIterator.hasNext(BufferedRowIterator.java:43)
at org.apache.spark.sql.execution.WholeStageCodegenExec$$anon$1.hasNext(WholeStageCodegenExec.scala:753)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$getByteArrayRdd$1(SparkPlan.scala:340)
at org.apache.spark.rdd.RDD.$anonfun$mapPartitionsInternal$2(RDD.scala:898)
at org.apache.spark.rdd.RDD.$anonfun$mapPartitionsInternal$2$adapted(RDD.scala:898)
at org.apache.spark.rdd.MapPartitionsRDD.compute(MapPartitionsRDD.scala:52)
at org.apache.spark.rdd.RDD.computeOrReadCheckpoint(RDD.scala:373)
at org.apache.spark.rdd.RDD.iterator(RDD.scala:337)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:127)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:469)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1377)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:472)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
... 1 more
```
</details>
<br/>
Another example with Python 3.7:
```python
sql("a")
```
<details>
<summary>Python exception message when <code>spark.sql.pyspark.jvmStacktrace.enabled</code> is off (default)</summary>
```
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "/.../spark/python/pyspark/sql/session.py", line 646, in sql
return DataFrame(self._jsparkSession.sql(sqlQuery), self._wrapped)
File "/.../spark/python/lib/py4j-0.10.9-src.zip/py4j/java_gateway.py", line 1305, in __call__
File "/.../spark/python/pyspark/sql/utils.py", line 131, in deco
raise_from(converted)
File "<string>", line 3, in raise_from
pyspark.sql.utils.ParseException:
mismatched input 'a' expecting {'(', 'ADD', 'ALTER', 'ANALYZE', 'CACHE', 'CLEAR', 'COMMENT', 'COMMIT', 'CREATE', 'DELETE', 'DESC', 'DESCRIBE', 'DFS', 'DROP', 'EXPLAIN', 'EXPORT', 'FROM', 'GRANT', 'IMPORT', 'INSERT', 'LIST', 'LOAD', 'LOCK', 'MAP', 'MERGE', 'MSCK', 'REDUCE', 'REFRESH', 'REPLACE', 'RESET', 'REVOKE', 'ROLLBACK', 'SELECT', 'SET', 'SHOW', 'START', 'TABLE', 'TRUNCATE', 'UNCACHE', 'UNLOCK', 'UPDATE', 'USE', 'VALUES', 'WITH'}(line 1, pos 0)
== SQL ==
a
^^^
```
</details>
<details>
<summary>Python exception message when <code>spark.sql.pyspark.jvmStacktrace.enabled</code> is on</summary>
```
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "/.../spark/python/pyspark/sql/session.py", line 646, in sql
return DataFrame(self._jsparkSession.sql(sqlQuery), self._wrapped)
File "/.../spark/python/lib/py4j-0.10.9-src.zip/py4j/java_gateway.py", line 1305, in __call__
File "/.../spark/python/pyspark/sql/utils.py", line 131, in deco
raise_from(converted)
File "<string>", line 3, in raise_from
pyspark.sql.utils.ParseException:
mismatched input 'a' expecting {'(', 'ADD', 'ALTER', 'ANALYZE', 'CACHE', 'CLEAR', 'COMMENT', 'COMMIT', 'CREATE', 'DELETE', 'DESC', 'DESCRIBE', 'DFS', 'DROP', 'EXPLAIN', 'EXPORT', 'FROM', 'GRANT', 'IMPORT', 'INSERT', 'LIST', 'LOAD', 'LOCK', 'MAP', 'MERGE', 'MSCK', 'REDUCE', 'REFRESH', 'REPLACE', 'RESET', 'REVOKE', 'ROLLBACK', 'SELECT', 'SET', 'SHOW', 'START', 'TABLE', 'TRUNCATE', 'UNCACHE', 'UNLOCK', 'UPDATE', 'USE', 'VALUES', 'WITH'}(line 1, pos 0)
== SQL ==
a
^^^
JVM stacktrace:
org.apache.spark.sql.catalyst.parser.ParseException:
mismatched input 'a' expecting {'(', 'ADD', 'ALTER', 'ANALYZE', 'CACHE', 'CLEAR', 'COMMENT', 'COMMIT', 'CREATE', 'DELETE', 'DESC', 'DESCRIBE', 'DFS', 'DROP', 'EXPLAIN', 'EXPORT', 'FROM', 'GRANT', 'IMPORT', 'INSERT', 'LIST', 'LOAD', 'LOCK', 'MAP', 'MERGE', 'MSCK', 'REDUCE', 'REFRESH', 'REPLACE', 'RESET', 'REVOKE', 'ROLLBACK', 'SELECT', 'SET', 'SHOW', 'START', 'TABLE', 'TRUNCATE', 'UNCACHE', 'UNLOCK', 'UPDATE', 'USE', 'VALUES', 'WITH'}(line 1, pos 0)
== SQL ==
a
^^^
at org.apache.spark.sql.catalyst.parser.ParseException.withCommand(ParseDriver.scala:266)
at org.apache.spark.sql.catalyst.parser.AbstractSqlParser.parse(ParseDriver.scala:133)
at org.apache.spark.sql.execution.SparkSqlParser.parse(SparkSqlParser.scala:49)
at org.apache.spark.sql.catalyst.parser.AbstractSqlParser.parsePlan(ParseDriver.scala:81)
at org.apache.spark.sql.SparkSession.$anonfun$sql$2(SparkSession.scala:604)
at org.apache.spark.sql.catalyst.QueryPlanningTracker.measurePhase(QueryPlanningTracker.scala:111)
at org.apache.spark.sql.SparkSession.$anonfun$sql$1(SparkSession.scala:604)
at org.apache.spark.sql.SparkSession.withActive(SparkSession.scala:763)
at org.apache.spark.sql.SparkSession.sql(SparkSession.scala:601)
at sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
at sun.reflect.NativeMethodAccessorImpl.invoke(NativeMethodAccessorImpl.java:62)
at sun.reflect.DelegatingMethodAccessorImpl.invoke(DelegatingMethodAccessorImpl.java:43)
at java.lang.reflect.Method.invoke(Method.java:498)
at py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
at py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:357)
at py4j.Gateway.invoke(Gateway.java:282)
at py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
at py4j.commands.CallCommand.execute(CallCommand.java:79)
at py4j.GatewayConnection.run(GatewayConnection.java:238)
at java.lang.Thread.run(Thread.java:748)
```
</details>
<details>
<summary>Python exception message without this change</summary>
```
Traceback (most recent call last):
File "/.../spark/python/pyspark/sql/utils.py", line 98, in deco
return f(*a, **kw)
File "/.../spark/python/lib/py4j-0.10.9-src.zip/py4j/protocol.py", line 328, in get_return_value
py4j.protocol.Py4JJavaError: An error occurred while calling o26.sql.
: org.apache.spark.sql.catalyst.parser.ParseException:
mismatched input 'a' expecting {'(', 'ADD', 'ALTER', 'ANALYZE', 'CACHE', 'CLEAR', 'COMMENT', 'COMMIT', 'CREATE', 'DELETE', 'DESC', 'DESCRIBE', 'DFS', 'DROP', 'EXPLAIN', 'EXPORT', 'FROM', 'GRANT', 'IMPORT', 'INSERT', 'LIST', 'LOAD', 'LOCK', 'MAP', 'MERGE', 'MSCK', 'REDUCE', 'REFRESH', 'REPLACE', 'RESET', 'REVOKE', 'ROLLBACK', 'SELECT', 'SET', 'SHOW', 'START', 'TABLE', 'TRUNCATE', 'UNCACHE', 'UNLOCK', 'UPDATE', 'USE', 'VALUES', 'WITH'}(line 1, pos 0)
== SQL ==
a
^^^
at org.apache.spark.sql.catalyst.parser.ParseException.withCommand(ParseDriver.scala:266)
at org.apache.spark.sql.catalyst.parser.AbstractSqlParser.parse(ParseDriver.scala:133)
at org.apache.spark.sql.execution.SparkSqlParser.parse(SparkSqlParser.scala:49)
at org.apache.spark.sql.catalyst.parser.AbstractSqlParser.parsePlan(ParseDriver.scala:81)
at org.apache.spark.sql.SparkSession.$anonfun$sql$2(SparkSession.scala:604)
at org.apache.spark.sql.catalyst.QueryPlanningTracker.measurePhase(QueryPlanningTracker.scala:111)
at org.apache.spark.sql.SparkSession.$anonfun$sql$1(SparkSession.scala:604)
at org.apache.spark.sql.SparkSession.withActive(SparkSession.scala:763)
at org.apache.spark.sql.SparkSession.sql(SparkSession.scala:601)
at sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
at sun.reflect.NativeMethodAccessorImpl.invoke(NativeMethodAccessorImpl.java:62)
at sun.reflect.DelegatingMethodAccessorImpl.invoke(DelegatingMethodAccessorImpl.java:43)
at java.lang.reflect.Method.invoke(Method.java:498)
at py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
at py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:357)
at py4j.Gateway.invoke(Gateway.java:282)
at py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
at py4j.commands.CallCommand.execute(CallCommand.java:79)
at py4j.GatewayConnection.run(GatewayConnection.java:238)
at java.lang.Thread.run(Thread.java:748)
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "/.../spark/python/pyspark/sql/session.py", line 646, in sql
return DataFrame(self._jsparkSession.sql(sqlQuery), self._wrapped)
File "/.../spark/python/lib/py4j-0.10.9-src.zip/py4j/java_gateway.py", line 1305, in __call__
File "/.../spark/python/pyspark/sql/utils.py", line 102, in deco
raise converted
pyspark.sql.utils.ParseException:
mismatched input 'a' expecting {'(', 'ADD', 'ALTER', 'ANALYZE', 'CACHE', 'CLEAR', 'COMMENT', 'COMMIT', 'CREATE', 'DELETE', 'DESC', 'DESCRIBE', 'DFS', 'DROP', 'EXPLAIN', 'EXPORT', 'FROM', 'GRANT', 'IMPORT', 'INSERT', 'LIST', 'LOAD', 'LOCK', 'MAP', 'MERGE', 'MSCK', 'REDUCE', 'REFRESH', 'REPLACE', 'RESET', 'REVOKE', 'ROLLBACK', 'SELECT', 'SET', 'SHOW', 'START', 'TABLE', 'TRUNCATE', 'UNCACHE', 'UNLOCK', 'UPDATE', 'USE', 'VALUES', 'WITH'}(line 1, pos 0)
== SQL ==
a
^^^
```
</details>
### Why are the changes needed?
Currently, PySpark exceptions are very unfriendly to Python users with causing a bunch of JVM stacktrace. See "Python exception message without this change" above.
### Does this PR introduce _any_ user-facing change?
Yes, it will change the exception message. See the examples above.
### How was this patch tested?
Manually tested by
```bash
./bin/pyspark --conf spark.sql.pyspark.jvmStacktrace.enabled=true
```
and running the examples above.
Closes#28661 from HyukjinKwon/python-debug.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR manually specifies the class for the input array being used in `(SparkContext|StreamingContext).union`. It fixes a regression introduced from SPARK-25737.
```python
rdd1 = sc.parallelize([1,2,3,4,5])
rdd2 = sc.parallelize([6,7,8,9,10])
pairRDD1 = rdd1.zip(rdd2)
sc.union([pairRDD1, pairRDD1]).collect()
```
in the current master and `branch-3.0`:
```
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "/.../spark/python/pyspark/context.py", line 870, in union
jrdds[i] = rdds[i]._jrdd
File "/.../spark/python/lib/py4j-0.10.9-src.zip/py4j/java_collections.py", line 238, in __setitem__
File "/.../spark/python/lib/py4j-0.10.9-src.zip/py4j/java_collections.py", line 221, in __set_item
File "/.../spark/python/lib/py4j-0.10.9-src.zip/py4j/protocol.py", line 332, in get_return_value
py4j.protocol.Py4JError: An error occurred while calling None.None. Trace:
py4j.Py4JException: Cannot convert org.apache.spark.api.java.JavaPairRDD to org.apache.spark.api.java.JavaRDD
at py4j.commands.ArrayCommand.convertArgument(ArrayCommand.java:166)
at py4j.commands.ArrayCommand.setArray(ArrayCommand.java:144)
at py4j.commands.ArrayCommand.execute(ArrayCommand.java:97)
at py4j.GatewayConnection.run(GatewayConnection.java:238)
at java.lang.Thread.run(Thread.java:748)
```
which works in Spark 2.4.5:
```
[(1, 6), (2, 7), (3, 8), (4, 9), (5, 10), (1, 6), (2, 7), (3, 8), (4, 9), (5, 10)]
```
It assumed the class of the input array is the same `JavaRDD` or `JavaDStream`; however, that can be different such as `JavaPairRDD`.
This fix is based on redsanket's initial approach, and will be co-authored.
### Why are the changes needed?
To fix a regression from Spark 2.4.5.
### Does this PR introduce _any_ user-facing change?
No, it's only in unreleased branches. This is to fix a regression.
### How was this patch tested?
Manually tested, and a unittest was added.
Closes#28648 from HyukjinKwon/SPARK-31788.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
Add instance weight support in LogisticRegressionSummary
### Why are the changes needed?
LogisticRegression, MulticlassClassificationEvaluator and BinaryClassificationEvaluator support instance weight. We should support instance weight in LogisticRegressionSummary too.
### Does this PR introduce _any_ user-facing change?
No
### How was this patch tested?
Add new tests
Closes#28657 from huaxingao/weighted_summary.
Authored-by: Huaxin Gao <huaxing@us.ibm.com>
Signed-off-by: Sean Owen <srowen@gmail.com>
### What changes were proposed in this pull request?
Followup to make assertions from recent test consistent with the rest of the module
### Why are the changes needed?
Better to use assertions from `unittest` and be consistent
### Does this PR introduce _any_ user-facing change?
No
### How was this patch tested?
Existing tests
Closes#28659 from BryanCutler/arrow-category-test-fix-SPARK-25351.
Authored-by: Bryan Cutler <cutlerb@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
Adds `inputFiles()` method to PySpark `DataFrame`. Using this, PySpark users can list all files constituting a `DataFrame`.
**Before changes:**
```
>>> spark.read.load("examples/src/main/resources/people.json", format="json").inputFiles()
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "/***/***/spark/python/pyspark/sql/dataframe.py", line 1388, in __getattr__
"'%s' object has no attribute '%s'" % (self.__class__.__name__, name))
AttributeError: 'DataFrame' object has no attribute 'inputFiles'
```
**After changes:**
```
>>> spark.read.load("examples/src/main/resources/people.json", format="json").inputFiles()
[u'file:///***/***/spark/examples/src/main/resources/people.json']
```
### Why are the changes needed?
This method is already supported for spark with scala and java.
### Does this PR introduce _any_ user-facing change?
Yes, Now users can list all files of a DataFrame using `inputFiles()`
### How was this patch tested?
UT added.
Closes#28652 from iRakson/SPARK-31763.
Authored-by: iRakson <raksonrakesh@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
Handle Pandas category type while converting from python with Arrow enabled. The category column will be converted to whatever type the category elements are as is the case with Arrow disabled.
### Does this PR introduce any user-facing change?
No
### How was this patch tested?
New unit tests were added for `createDataFrame` and scalar `pandas_udf`
Closes#26585 from jalpan-randeri/feature-pyarrow-dictionary-type.
Authored-by: Jalpan Randeri <randerij@amazon.com>
Signed-off-by: Bryan Cutler <cutlerb@gmail.com>
### What changes were proposed in this pull request?
Add weight support in ClusteringEvaluator
### Why are the changes needed?
Currently, BinaryClassificationEvaluator, RegressionEvaluator, and MulticlassClassificationEvaluator support instance weight, but ClusteringEvaluator doesn't, so we will add instance weight support in ClusteringEvaluator.
### Does this PR introduce _any_ user-facing change?
Yes.
ClusteringEvaluator.setWeightCol
### How was this patch tested?
add new unit test
Closes#28553 from huaxingao/weight_evaluator.
Authored-by: Huaxin Gao <huaxing@us.ibm.com>
Signed-off-by: Sean Owen <srowen@gmail.com>
### What changes were proposed in this pull request?
UnionRDD of PairRDDs causing a bug. The fix is to check for instance type before proceeding
### Why are the changes needed?
Changes are needed to avoid users running into issues with union rdd operation with any other type other than JavaRDD.
### Does this PR introduce _any_ user-facing change?
Yes
Before:
SparkSession available as 'spark'.
>>> rdd1 = sc.parallelize([1,2,3,4,5])
>>> rdd2 = sc.parallelize([6,7,8,9,10])
>>> pairRDD1 = rdd1.zip(rdd2)
>>> unionRDD1 = sc.union([pairRDD1, pairRDD1])
Traceback (most recent call last): File "<stdin>", line 1, in <module> File "/home/gs/spark/latest/python/pyspark/context.py", line 870,
in union jrdds[i] = rdds[i]._jrdd
File "/home/gs/spark/latest/python/lib/py4j-0.10.9-src.zip/py4j/java_collections.py", line 238, in setitem File "/home/gs/spark/latest/python/lib/py4j-0.10.9-src.zip/py4j/java_collections.py", line 221,
in __set_item File "/home/gs/spark/latest/python/lib/py4j-0.10.9-src.zip/py4j/protocol.py", line 332, in get_return_value py4j.protocol.Py4JError: An error occurred while calling None.None. Trace: py4j.Py4JException: Cannot convert org.apache.spark.api.java.JavaPairRDD to org.apache.spark.api.java.JavaRDD at py4j.commands.ArrayCommand.convertArgument(ArrayCommand.java:166) at py4j.commands.ArrayCommand.setArray(ArrayCommand.java:144) at py4j.commands.ArrayCommand.execute(ArrayCommand.java:97) at py4j.GatewayConnection.run(GatewayConnection.java:238) at java.lang.Thread.run(Thread.java:748)
After:
>>> rdd2 = sc.parallelize([6,7,8,9,10])
>>> pairRDD1 = rdd1.zip(rdd2)
>>> unionRDD1 = sc.union([pairRDD1, pairRDD1])
>>> unionRDD1.collect()
[(1, 6), (2, 7), (3, 8), (4, 9), (5, 10), (1, 6), (2, 7), (3, 8), (4, 9), (5, 10)]
### How was this patch tested?
Tested with the reproduced piece of code above manually
Closes#28603 from redsanket/SPARK-31788.
Authored-by: schintap <schintap@verizonmedia.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR proposes to only allow the import of `ResourceInformation` as below:
```
pyspark.resource.ResourceInformation
```
instead of
```
pyspark.ResourceInformation
pyspark.resource.ResourceInformation
```
because `pyspark.resource` is a separate module, and it is documented so.
The constructor of `ResourceInformation` isn't supposed to directly call anyway.
### Why are the changes needed?
To keep the code structure coherent.
### Does this PR introduce _any_ user-facing change?
No, it will be in the unreleased branches.
### How was this patch tested?
Manually tested via importing:
Before:
```python
>>> import pyspark
>>> pyspark.ResourceInformation
<class 'pyspark.resource.information.ResourceInformation'>
>>> pyspark.resource.ResourceInformation
<class 'pyspark.resource.information.ResourceInformation'>
```
After:
```python
>>> import pyspark
>>> pyspark.ResourceInformation
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
AttributeError: module 'pyspark' has no attribute 'ResourceInformation'
>>> pyspark.resource.ResourceInformation
<class 'pyspark.resource.information.ResourceInformation'>
```
Also tested via
```bash
cd python
./run-tests --python-executables=python3 --modules=pyspark-core,pyspark-resource
```
Jenkins will test and existing tests should cover.
Closes#28589 from HyukjinKwon/SPARK-31767.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: Dongjoon Hyun <dongjoon@apache.org>
### What changes were proposed in this pull request?
This PR is kind of a followup for SPARK-29641 and SPARK-28234. This PR proposes:
1.. Document the new `pyspark.resource` module introduced at 95aec091e4, in PySpark API docs.
2.. Move classes into fewer and simpler modules
Before:
```
pyspark
├── resource
│ ├── executorrequests.py
│ │ ├── class ExecutorResourceRequest
│ │ └── class ExecutorResourceRequests
│ ├── taskrequests.py
│ │ ├── class TaskResourceRequest
│ │ └── class TaskResourceRequests
│ ├── resourceprofilebuilder.py
│ │ └── class ResourceProfileBuilder
│ ├── resourceprofile.py
│ │ └── class ResourceProfile
└── resourceinformation
└── class ResourceInformation
```
After:
```
pyspark
└── resource
├── requests.py
│ ├── class ExecutorResourceRequest
│ ├── class ExecutorResourceRequests
│ ├── class TaskResourceRequest
│ └── class TaskResourceRequests
├── profile.py
│ ├── class ResourceProfileBuilder
│ └── class ResourceProfile
└── information.py
└── class ResourceInformation
```
3.. Minor docstring fix e.g.:
```diff
- param name the name of the resource
- param addresses an array of strings describing the addresses of the resource
+ :param name: the name of the resource
+ :param addresses: an array of strings describing the addresses of the resource
+
+ .. versionadded:: 3.0.0
```
### Why are the changes needed?
To document APIs, and move Python modules to fewer and simpler modules.
### Does this PR introduce _any_ user-facing change?
No, the changes are in unreleased branches.
### How was this patch tested?
Manually tested via:
```bash
cd python
./run-tests --python-executables=python3 --modules=pyspark-core
./run-tests --python-executables=python3 --modules=pyspark-resource
```
Closes#28569 from HyukjinKwon/SPARK-28234-SPARK-29641-followup.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: Dongjoon Hyun <dongjoon@apache.org>
This commit is published into the public domain.
### What changes were proposed in this pull request?
Some syntax issues in docstrings have been fixed.
### Why are the changes needed?
In some places, the documentation did not render as intended, e.g. parameter documentations were not formatted as such.
### Does this PR introduce any user-facing change?
Slight improvements in documentation.
### How was this patch tested?
Manual testing and `dev/lint-python` run. No new Sphinx warnings arise due to this change.
Closes#28559 from DavidToneian/SPARK-31739.
Authored-by: David Toneian <david@toneian.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
Return LogisticRegressionSummary for multiclass logistic regression evaluate in PySpark
### Why are the changes needed?
Currently we have
```
since("2.0.0")
def evaluate(self, dataset):
if not isinstance(dataset, DataFrame):
raise ValueError("dataset must be a DataFrame but got %s." % type(dataset))
java_blr_summary = self._call_java("evaluate", dataset)
return BinaryLogisticRegressionSummary(java_blr_summary)
```
we should return LogisticRegressionSummary for multiclass logistic regression
### Does this PR introduce _any_ user-facing change?
Yes
return LogisticRegressionSummary instead of BinaryLogisticRegressionSummary for multiclass logistic regression in Python
### How was this patch tested?
unit test
Closes#28503 from huaxingao/lr_summary.
Authored-by: Huaxin Gao <huaxing@us.ibm.com>
Signed-off-by: Sean Owen <srowen@gmail.com>
### What changes were proposed in this pull request?
1, add new param blockSize;
2, if blockSize==1, keep original behavior, code path trainOnRows;
3, if blockSize>1, standardize and stack input vectors to blocks (like ALS/MLP), code path trainOnBlocks
### Why are the changes needed?
performance gain on dense dataset HIGGS:
1, save about 45% RAM;
2, 3X faster with openBLAS
### Does this PR introduce any user-facing change?
add a new expert param `blockSize`
### How was this patch tested?
added testsuites
Closes#27473 from zhengruifeng/blockify_gmm.
Authored-by: zhengruifeng <ruifengz@foxmail.com>
Signed-off-by: zhengruifeng <ruifengz@foxmail.com>
### What changes were proposed in this pull request?
Add Python version of
```
Since("3.1.0")
def test(
dataset: DataFrame,
featuresCol: String,
labelCol: String,
flatten: Boolean): DataFrame
```
### Why are the changes needed?
parity between scala and python
### Does this PR introduce _any_ user-facing change?
yes
new method
```
Since("3.1.0")
def test(
dataset: DataFrame,
featuresCol: String,
labelCol: String,
flatten: Boolean): DataFrame
```
in PySpark ANOVATest/ChisqTest/FValueTest
### How was this patch tested?
New doctest
Closes#28483 from huaxingao/flatten_py.
Authored-by: Huaxin Gao <huaxing@us.ibm.com>
Signed-off-by: Sean Owen <srowen@gmail.com>
### What changes were proposed in this pull request?
1, add new param blockSize;
2, add a new class InstanceBlock;
3, if blockSize==1, keep original behavior; if blockSize>1, stack input vectors to blocks (like ALS/MLP);
4, if blockSize>1, standardize the input outside of optimization procedure;
### Why are the changes needed?
it will obtain performance gain on dense datasets, such as epsilon
1, reduce RAM to persist traing dataset; (save about 40% RAM)
2, use Level-2 BLAS routines; (~10X speedup)
### Does this PR introduce _any_ user-facing change?
Yes, a new param is added
### How was this patch tested?
existing and added testsuites
Closes#28473 from zhengruifeng/blockify_aft.
Authored-by: zhengruifeng <ruifengz@foxmail.com>
Signed-off-by: zhengruifeng <ruifengz@foxmail.com>
### What changes were proposed in this pull request?
Add ANOVASelector and FValueSelector to PySpark
### Why are the changes needed?
ANOVASelector and FValueSelector have been implemented in Scala. We need to implement these in Python as well.
### Does this PR introduce _any_ user-facing change?
Yes. Add Python version of ANOVASelector and FValueSelector
### How was this patch tested?
new doctest
Closes#28464 from huaxingao/selector_py.
Authored-by: Huaxin Gao <huaxing@us.ibm.com>
Signed-off-by: zhengruifeng <ruifengz@foxmail.com>
### What changes were proposed in this pull request?
1, add new param blockSize;
2, add a new class InstanceBlock;
3, if blockSize==1, keep original behavior; if blockSize>1, stack input vectors to blocks (like ALS/MLP);
4, if blockSize>1, standardize the input outside of optimization procedure;
### Why are the changes needed?
it will obtain performance gain on dense datasets, such as `epsilon`
1, reduce RAM to persist traing dataset; (save about 40% RAM)
2, use Level-2 BLAS routines; (up to 6X(squaredError)~12X(huber) speedup)
### Does this PR introduce _any_ user-facing change?
Yes, a new param is added
### How was this patch tested?
existing and added testsuites
Closes#28471 from zhengruifeng/blockify_lir_II.
Authored-by: zhengruifeng <ruifengz@foxmail.com>
Signed-off-by: zhengruifeng <ruifengz@foxmail.com>
### What changes were proposed in this pull request?
The RuntimeReplaceable ones are runtime replaceable, thus, their original parameters are not going to be resolved to PrettyAttribute and remain debug style string if we directly implement their `sql` methods with their parameters' `sql` methods.
This PR is raised with suggestions by maropu and cloud-fan https://github.com/apache/spark/pull/28402/files#r417656589. In this PR, we re-implement the `sql` methods of the RuntimeReplaceable ones with toPettySQL
### Why are the changes needed?
Consistency of schema output between RuntimeReplaceable expressions and normal ones.
For example, `date_format` vs `to_timestamp`, before this PR, they output differently
#### Before
```sql
select date_format(timestamp '2019-10-06', 'yyyy-MM-dd uuuu')
struct<date_format(TIMESTAMP '2019-10-06 00:00:00', yyyy-MM-dd uuuu):string>
select to_timestamp("2019-10-06S10:11:12.12345", "yyyy-MM-dd'S'HH:mm:ss.SSSSSS")
struct<to_timestamp('2019-10-06S10:11:12.12345', 'yyyy-MM-dd\'S\'HH:mm:ss.SSSSSS'):timestamp>
```
#### After
```sql
select date_format(timestamp '2019-10-06', 'yyyy-MM-dd uuuu')
struct<date_format(TIMESTAMP '2019-10-06 00:00:00', yyyy-MM-dd uuuu):string>
select to_timestamp("2019-10-06T10:11:12'12", "yyyy-MM-dd'T'HH:mm:ss''SSSS")
struct<to_timestamp(2019-10-06T10:11:12'12, yyyy-MM-dd'T'HH:mm:ss''SSSS):timestamp>
````
### Does this PR introduce _any_ user-facing change?
Yes, the schema output style changed for the runtime replaceable expressions as shown in the above example
### How was this patch tested?
regenerate all related tests
Closes#28420 from yaooqinn/SPARK-31615.
Authored-by: Kent Yao <yaooqinn@hotmail.com>
Signed-off-by: Takeshi Yamamuro <yamamuro@apache.org>
### What changes were proposed in this pull request?
1, reorg the `fit` method in LR to several blocks (`createModel`, `createBounds`, `createOptimizer`, `createInitCoefWithInterceptMatrix`);
2, add new param blockSize;
3, if blockSize==1, keep original behavior, code path `trainOnRows`;
4, if blockSize>1, standardize and stack input vectors to blocks (like ALS/MLP), code path `trainOnBlocks`
### Why are the changes needed?
On dense dataset `epsilon_normalized.t`:
1, reduce RAM to persist traing dataset; (save about 40% RAM)
2, use Level-2 BLAS routines; (4x ~ 5x faster)
### Does this PR introduce _any_ user-facing change?
Yes, a new param is added
### How was this patch tested?
existing and added testsuites
Closes#28458 from zhengruifeng/blockify_lor_II.
Authored-by: zhengruifeng <ruifengz@foxmail.com>
Signed-off-by: zhengruifeng <ruifengz@foxmail.com>
### What changes were proposed in this pull request?
Add VarianceThresholdSelector to PySpark
### Why are the changes needed?
parity between Scala and Python
### Does this PR introduce any user-facing change?
Yes.
VarianceThresholdSelector is added to PySpark
### How was this patch tested?
new doctest
Closes#28409 from huaxingao/variance_py.
Authored-by: Huaxin Gao <huaxing@us.ibm.com>
Signed-off-by: Sean Owen <srowen@gmail.com>
### What changes were proposed in this pull request?
1, add new param `blockSize`;
2, add a new class InstanceBlock;
3, **if `blockSize==1`, keep original behavior; if `blockSize>1`, stack input vectors to blocks (like ALS/MLP);**
4, if `blockSize>1`, standardize the input outside of optimization procedure;
### Why are the changes needed?
1, reduce RAM to persist traing dataset; (save about 40% RAM)
2, use Level-2 BLAS routines; (4x ~ 5x faster on dataset `epsilon`)
### Does this PR introduce any user-facing change?
Yes, a new param is added
### How was this patch tested?
existing and added testsuites
Closes#28349 from zhengruifeng/blockify_svc_II.
Authored-by: zhengruifeng <ruifengz@foxmail.com>
Signed-off-by: zhengruifeng <ruifengz@foxmail.com>
### What changes were proposed in this pull request?
I add a new API in pyspark RDD class:
def collectWithJobGroup(self, groupId, description, interruptOnCancel=False)
This API do the same thing with `rdd.collect`, but it can specify the job group when do collect.
The purpose of adding this API is, if we use:
```
sc.setJobGroup("group-id...")
rdd.collect()
```
The `setJobGroup` API in pyspark won't work correctly. This related to a bug discussed in
https://issues.apache.org/jira/browse/SPARK-31549
Note:
This PR is a rather temporary workaround for `PYSPARK_PIN_THREAD`, and as a step to migrate to `PYSPARK_PIN_THREAD` smoothly. It targets Spark 3.0.
- `PYSPARK_PIN_THREAD` is unstable at this moment that affects whole PySpark applications.
- It is impossible to make it runtime configuration as it has to be set before JVM is launched.
- There is a thread leak issue between Python and JVM. We should address but it's not a release blocker for Spark 3.0 since the feature is experimental. I plan to handle this after Spark 3.0 due to stability.
Once `PYSPARK_PIN_THREAD` is enabled by default, we should remove this API out ideally. I will target to deprecate this API in Spark 3.1.
### Why are the changes needed?
Fix bug.
### Does this PR introduce any user-facing change?
A develop API in pyspark: `pyspark.RDD. collectWithJobGroup`
### How was this patch tested?
Unit test.
Closes#28395 from WeichenXu123/collect_with_job_group.
Authored-by: Weichen Xu <weichen.xu@databricks.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR proposes to use a different approach instead of breaking it per Micheal's rubric added at https://spark.apache.org/versioning-policy.html. It deprecates the behaviour for now. It will be gradually removed in the future releases.
After this change,
```python
import warnings
warnings.simplefilter("always")
from pyspark.sql.functions import *
df = spark.range(2)
map_col = create_map(lit(0), lit(100), lit(1), lit(200))
df.withColumn("mapped", map_col.getItem(col('id'))).show()
```
```
/.../python/pyspark/sql/column.py:311: DeprecationWarning: A column as 'key' in getItem is
deprecated as of Spark 3.0, and will not be supported in the future release. Use `column[key]`
or `column.key` syntax instead.
DeprecationWarning)
...
```
```python
import warnings
warnings.simplefilter("always")
from pyspark.sql.functions import *
df = spark.range(2)
struct_col = struct(lit(0), lit(100), lit(1), lit(200))
df.withColumn("struct", struct_col.getField(lit("col1"))).show()
```
```
/.../spark/python/pyspark/sql/column.py:336: DeprecationWarning: A column as 'name'
in getField is deprecated as of Spark 3.0, and will not be supported in the future release. Use
`column[name]` or `column.name` syntax instead.
DeprecationWarning)
```
### Why are the changes needed?
To prevent the radical behaviour change after the amended versioning policy.
### Does this PR introduce any user-facing change?
Yes, it will show the deprecated warning message.
### How was this patch tested?
Manually tested.
Closes#28327 from HyukjinKwon/SPARK-29664.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
Fix Pyspark CrossValidator/TrainValidationSplit with pipeline estimator cannot save and load model.
Most pyspark estimators/transformers inherit `JavaParams`, but some estimators are special (in order to support pure python implemented nested estimators/transformers):
* Pipeline
* OneVsRest
* CrossValidator
* TrainValidationSplit
But note that, currently, in pyspark, estimators listed above, their model reader/writer do NOT support pure python implemented nested estimators/transformers. Because they use java reader/writer wrapper as python side reader/writer.
Pyspark CrossValidator/TrainValidationSplit model reader/writer require all estimators define the `_transfer_param_map_to_java` and `_transfer_param_map_from_java` (used in model read/write).
OneVsRest class already defines the two methods, but Pipeline do not, so it lead to this bug.
In this PR I add `_transfer_param_map_to_java` and `_transfer_param_map_from_java` into Pipeline class.
### Why are the changes needed?
Bug fix.
### Does this PR introduce any user-facing change?
No
### How was this patch tested?
Unit test.
Manually test in pyspark shell:
1) CrossValidator with Simple Pipeline estimator
```
from pyspark.ml import Pipeline
from pyspark.ml.classification import LogisticRegression
from pyspark.ml.evaluation import BinaryClassificationEvaluator
from pyspark.ml.feature import HashingTF, Tokenizer
from pyspark.ml.tuning import CrossValidator, CrossValidatorModel, ParamGridBuilder
training = spark.createDataFrame([
(0, "a b c d e spark", 1.0),
(1, "b d", 0.0),
(2, "spark f g h", 1.0),
(3, "hadoop mapreduce", 0.0),
(4, "b spark who", 1.0),
(5, "g d a y", 0.0),
(6, "spark fly", 1.0),
(7, "was mapreduce", 0.0),
], ["id", "text", "label"])
# Configure an ML pipeline, which consists of tree stages: tokenizer, hashingTF, and lr.
tokenizer = Tokenizer(inputCol="text", outputCol="words")
hashingTF = HashingTF(inputCol=tokenizer.getOutputCol(), outputCol="features")
lr = LogisticRegression(maxIter=10)
pipeline = Pipeline(stages=[tokenizer, hashingTF, lr])
paramGrid = ParamGridBuilder() \
.addGrid(hashingTF.numFeatures, [10, 100, 1000]) \
.addGrid(lr.regParam, [0.1, 0.01]) \
.build()
crossval = CrossValidator(estimator=pipeline,
estimatorParamMaps=paramGrid,
evaluator=BinaryClassificationEvaluator(),
numFolds=2) # use 3+ folds in practice
# Run cross-validation, and choose the best set of parameters.
cvModel = crossval.fit(training)
cvModel.save('/tmp/cv_model001')
CrossValidatorModel.load('/tmp/cv_model001')
```
2) CrossValidator with Pipeline estimator which include a OneVsRest estimator stage, and OneVsRest estimator nest a LogisticRegression estimator.
```
from pyspark.ml.linalg import Vectors
from pyspark.ml import Estimator, Model
from pyspark.ml.classification import LogisticRegression, LogisticRegressionModel, OneVsRest
from pyspark.ml.evaluation import BinaryClassificationEvaluator, \
MulticlassClassificationEvaluator, RegressionEvaluator
from pyspark.ml.linalg import Vectors
from pyspark.ml.param import Param, Params
from pyspark.ml.tuning import CrossValidator, CrossValidatorModel, ParamGridBuilder, \
TrainValidationSplit, TrainValidationSplitModel
from pyspark.sql.functions import rand
from pyspark.testing.mlutils import SparkSessionTestCase
dataset = spark.createDataFrame(
[(Vectors.dense([0.0]), 0.0),
(Vectors.dense([0.4]), 1.0),
(Vectors.dense([0.5]), 0.0),
(Vectors.dense([0.6]), 1.0),
(Vectors.dense([1.0]), 1.0)] * 10,
["features", "label"])
ova = OneVsRest(classifier=LogisticRegression())
lr1 = LogisticRegression().setMaxIter(100)
lr2 = LogisticRegression().setMaxIter(150)
grid = ParamGridBuilder().addGrid(ova.classifier, [lr1, lr2]).build()
evaluator = MulticlassClassificationEvaluator()
pipeline = Pipeline(stages=[ova])
cv = CrossValidator(estimator=pipeline, estimatorParamMaps=grid, evaluator=evaluator)
cvModel = cv.fit(dataset)
cvModel.save('/tmp/model002')
cvModel2 = CrossValidatorModel.load('/tmp/model002')
```
TrainValidationSplit testing code are similar so I do not paste them.
Closes#28279 from WeichenXu123/fix_pipeline_tuning.
Authored-by: Weichen Xu <weichen.xu@databricks.com>
Signed-off-by: Xiangrui Meng <meng@databricks.com>
### What changes were proposed in this pull request?
As part of the Stage level scheduling features, add the Python api's to set resource profiles.
This also adds the functionality to properly apply the pyspark memory configuration when specified in the ResourceProfile. The pyspark memory configuration is being passed in the task local properties. This was an easy way to get it to the PythonRunner that needs it. I modeled this off how the barrier task scheduling is passing the addresses. As part of this I added in the JavaRDD api's because those are needed by python.
### Why are the changes needed?
python api for this feature
### Does this PR introduce any user-facing change?
Yes adds the java and python apis for user to specify a ResourceProfile to use stage level scheduling.
### How was this patch tested?
unit tests and manually tested on yarn. Tests also run to verify it errors properly on standalone and local mode where its not yet supported.
Closes#28085 from tgravescs/SPARK-29641-pr-base.
Lead-authored-by: Thomas Graves <tgraves@nvidia.com>
Co-authored-by: Thomas Graves <tgraves@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
1. Combine `BarrierRequestToSync` and `AllGatherRequestToSync` into `RequestToSync`, which is distinguished by `RequestMethod` type.
2. Remove unnecessary Json serialization/deserialization
3. Clean up some codes to make runBarrier() and `BarrierCoordinator` more general
4. Remove unused imports.
### Why are the changes needed?
To make codes simpler for better maintain in the future.
### Does this PR introduce any user-facing change?
No.
### How was this patch tested?
This is pure code refactor, so should be covered by existed tests.
Closes#28117 from Ngone51/refactor_barrier.
Authored-by: yi.wu <yi.wu@databricks.com>
Signed-off-by: Xingbo Jiang <xingbo.jiang@databricks.com>
### What changes were proposed in this pull request?
This PR is adding support duplicated column names for `toPandas` with Arrow execution.
### Why are the changes needed?
When we execute `toPandas()` with Arrow execution, it fails if the column names have duplicates.
```py
>>> spark.sql("select 1 v, 1 v").toPandas()
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "/path/to/lib/python3.7/site-packages/pyspark/sql/dataframe.py", line 2132, in toPandas
pdf = table.to_pandas()
File "pyarrow/array.pxi", line 441, in pyarrow.lib._PandasConvertible.to_pandas
File "pyarrow/table.pxi", line 1367, in pyarrow.lib.Table._to_pandas
File "/path/to/lib/python3.7/site-packages/pyarrow/pandas_compat.py", line 653, in table_to_blockmanager
columns = _deserialize_column_index(table, all_columns, column_indexes)
File "/path/to/lib/python3.7/site-packages/pyarrow/pandas_compat.py", line 704, in _deserialize_column_index
columns = _flatten_single_level_multiindex(columns)
File "/path/to/lib/python3.7/site-packages/pyarrow/pandas_compat.py", line 937, in _flatten_single_level_multiindex
raise ValueError('Found non-unique column index')
ValueError: Found non-unique column index
```
### Does this PR introduce any user-facing change?
Yes, previously we will face an error above, but after this PR, we will see the result:
```py
>>> spark.sql("select 1 v, 1 v").toPandas()
v v
0 1 1
```
### How was this patch tested?
Added and modified related tests.
Closes#28210 from ueshin/issues/SPARK-31441/to_pandas.
Authored-by: Takuya UESHIN <ueshin@databricks.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
Update default datetime pattern from `yyyy-MM-dd'T'HH:mm:ss.SSSXXX ` to `yyyy-MM-dd'T'HH:mm:ss[.SSS][XXX] ` for JSON/CSV APIs documentations
### Why are the changes needed?
doc fix
### Does this PR introduce any user-facing change?
Yes, the documentation will change
### How was this patch tested?
Passing Jenkins
Closes#28204 from yaooqinn/SPARK-31414-F.
Authored-by: Kent Yao <yaooqinn@hotmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR explicitly mention that the requirement of Iterator of Series to Iterator of Series and Iterator of Multiple Series to Iterator of Series (previously Scalar Iterator pandas UDF).
The actual limitation of this UDF is the same length of the _entire input and output_, instead of each series's length. Namely you can do something as below:
```python
from typing import Iterator, Tuple
import pandas as pd
from pyspark.sql.functions import pandas_udf
pandas_udf("long")
def func(
iterator: Iterator[pd.Series]) -> Iterator[pd.Series]:
return iter([pd.concat(iterator)])
spark.range(100).select(func("id")).show()
```
This characteristic allows you to prefetch the data from the iterator to speed up, compared to the regular Scalar to Scalar (previously Scalar pandas UDF).
### Why are the changes needed?
To document the correct restriction and characteristics of a feature.
### Does this PR introduce any user-facing change?
Yes in the documentation but only in unreleased branches.
### How was this patch tested?
Github Actions should test the documentation build
Closes#28160 from HyukjinKwon/SPARK-30722-followup.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR proposes to improve the error message from Scalar iterator pandas UDF.
### Why are the changes needed?
To show the correct error messages.
### Does this PR introduce any user-facing change?
Yes, but only in unreleased branches.
```python
import pandas as pd
from pyspark.sql.functions import pandas_udf, PandasUDFType
pandas_udf('long', PandasUDFType.SCALAR_ITER)
def pandas_plus_one(iterator):
for _ in iterator:
yield pd.Series(1)
spark.range(10).repartition(1).select(pandas_plus_one("id")).show()
```
```python
import pandas as pd
from pyspark.sql.functions import pandas_udf, PandasUDFType
pandas_udf('long', PandasUDFType.SCALAR_ITER)
def pandas_plus_one(iterator):
for _ in iterator:
yield pd.Series(list(range(20)))
spark.range(10).repartition(1).select(pandas_plus_one("id")).show()
```
**Before:**
```
RuntimeError: The number of output rows of pandas iterator UDF should
be the same with input rows. The input rows number is 10 but the output
rows number is 1.
```
```
AssertionError: Pandas MAP_ITER UDF outputted more rows than input rows.
```
**After:**
```
RuntimeError: The length of output in Scalar iterator pandas UDF should be
the same with the input's; however, the length of output was 1 and the length
of input was 10.
```
```
AssertionError: Pandas SCALAR_ITER UDF outputted more rows than input rows.
```
### How was this patch tested?
Unittests were fixed accordingly.
Closes#28135 from HyukjinKwon/SPARK-26412-followup.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR proposes to show a better error message when a user mistakenly installs `pyspark` from PIP but the default `python` does not point out the corresponding `pip`. See https://stackoverflow.com/questions/46286436/running-pyspark-after-pip-install-pyspark/49587560 as an example.
It can be reproduced as below:
I have two Python executables. `python` is Python 3.7, `pip` binds with Python 3.7 and `python2.7` is Python 2.7.
```bash
pip install pyspark
```
```bash
pyspark
```
```
...
Welcome to
____ __
/ __/__ ___ _____/ /__
_\ \/ _ \/ _ `/ __/ '_/
/__ / .__/\_,_/_/ /_/\_\ version 2.4.5
/_/
Using Python version 3.7.3 (default, Mar 27 2019 09:23:15)
SparkSession available as 'spark'.
...
```
```bash
PYSPARK_PYTHON=python2.7 pyspark
```
```
Could not find valid SPARK_HOME while searching ['/Users', '/usr/local/Cellar/python/3.7.5/Frameworks/Python.framework/Versions/3.7/bin']
/usr/local/Cellar/python/3.7.5/Frameworks/Python.framework/Versions/3.7/bin/pyspark: line 24: /bin/load-spark-env.sh: No such file or directory
/usr/local/Cellar/python/3.7.5/Frameworks/Python.framework/Versions/3.7/bin/pyspark: line 77: /bin/spark-submit: No such file or directory
/usr/local/Cellar/python/3.7.5/Frameworks/Python.framework/Versions/3.7/bin/pyspark: line 77: exec: /bin/spark-submit: cannot execute: No such file or directory
```
### Why are the changes needed?
There are multiple questions outside about this error and they have no idea what's going on. See:
- https://stackoverflow.com/questions/46286436/running-pyspark-after-pip-install-pyspark/49587560
- https://stackoverflow.com/questions/45991888/path-issue-could-not-find-valid-spark-home-while-searching
- https://stackoverflow.com/questions/49707239/pyspark-could-not-find-valid-spark-home
- https://stackoverflow.com/questions/55569985/pyspark-could-not-find-valid-spark-home
- https://stackoverflow.com/questions/48296474/error-could-not-find-valid-spark-home-while-searching-pycharm-in-windows
- https://github.com/ContinuumIO/anaconda-issues/issues/8076
The answer is usually setting `SPARK_HOME`; however this isn't completely correct.
It works if you set `SPARK_HOME` because `pyspark` executable script directly imports the library by using `SPARK_HOME` (see https://github.com/apache/spark/blob/master/bin/pyspark#L52-L53) instead of the default package location specified via `python` executable. So, this way you use a package installed in a different Python, which isn't ideal.
### Does this PR introduce any user-facing change?
Yes, it fixes the error message better.
**Before:**
```
Could not find valid SPARK_HOME while searching ['/Users', '/usr/local/Cellar/python/3.7.5/Frameworks/Python.framework/Versions/3.7/bin']
...
```
**After:**
```
Could not find valid SPARK_HOME while searching ['/Users', '/usr/local/Cellar/python/3.7.5/Frameworks/Python.framework/Versions/3.7/bin']
Did you install PySpark via a package manager such as pip or Conda? If so,
PySpark was not found in your Python environment. It is possible your
Python environment does not properly bind with your package manager.
Please check your default 'python' and if you set PYSPARK_PYTHON and/or
PYSPARK_DRIVER_PYTHON environment variables, and see if you can import
PySpark, for example, 'python -c 'import pyspark'.
If you cannot import, you can install by using the Python executable directly,
for example, 'python -m pip install pyspark [--user]'. Otherwise, you can also
explicitly set the Python executable, that has PySpark installed, to
PYSPARK_PYTHON or PYSPARK_DRIVER_PYTHON environment variables, for example,
'PYSPARK_PYTHON=python3 pyspark'.
...
```
### How was this patch tested?
Manually tested as described above.
Closes#28152 from HyukjinKwon/SPARK-31382.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This patch proposed to skip predicates on PythonUDFs to be pushdown through Aggregate.
### Why are the changes needed?
The predicates on PythonUDFs cannot be pushdown through Aggregate. Pushed down predicates cannot be evaluate because PythonUDFs cannot be evaluated on Filter and cause error like:
```
Caused by: java.lang.UnsupportedOperationException: Cannot generate code for expression: mean(input[1, struct<bar:bigint>, true].bar)
at org.apache.spark.sql.catalyst.expressions.Unevaluable.doGenCode(Expression.scala:304)
at org.apache.spark.sql.catalyst.expressions.Unevaluable.doGenCode$(Expression.scala:303)
at org.apache.spark.sql.catalyst.expressions.PythonUDF.doGenCode(PythonUDF.scala:52)
at org.apache.spark.sql.catalyst.expressions.Expression.$anonfun$genCode$3(Expression.scala:146)
at scala.Option.getOrElse(Option.scala:189)
at org.apache.spark.sql.catalyst.expressions.Expression.genCode(Expression.scala:141)
at org.apache.spark.sql.catalyst.expressions.CastBase.doGenCode(Cast.scala:821)
at org.apache.spark.sql.catalyst.expressions.Expression.$anonfun$genCode$3(Expression.scala:146)
at scala.Option.getOrElse(Option.scala:189)
```
### Does this PR introduce any user-facing change?
Yes. Previously the predicates on PythonUDFs will be pushdown through Aggregate can cause error. After this change, the query can work.
### How was this patch tested?
Unit test.
Closes#28089 from viirya/SPARK-30921.
Authored-by: Liang-Chi Hsieh <viirya@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
A small documentation change to clarify that the `rand()` function produces values in `[0.0, 1.0)`.
### Why are the changes needed?
`rand()` uses `Rand()` - which generates values in [0, 1) ([documented here](a1dbcd13a3/sql/catalyst/src/main/scala/org/apache/spark/sql/catalyst/expressions/randomExpressions.scala (L71))). The existing documentation suggests that 1.0 is a possible value returned by rand (i.e for a distribution written as `X ~ U(a, b)`, x can be a or b, so `U[0.0, 1.0]` suggests the value returned could include 1.0).
### Does this PR introduce any user-facing change?
Only documentation changes.
### How was this patch tested?
Documentation changes only.
Closes#28071 from Smeb/master.
Authored-by: Ben Ryves <benjamin.ryves@getyourguide.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
In the PR, I propose to update the doc for the `timeZone` option in JSON/CSV datasources and for the `tz` parameter of the `from_utc_timestamp()`/`to_utc_timestamp()` functions, and to restrict format of config's values to 2 forms:
1. Geographical regions, such as `America/Los_Angeles`.
2. Fixed offsets - a fully resolved offset from UTC. For example, `-08:00`.
### Why are the changes needed?
Other formats such as three-letter time zone IDs are ambitious, and depend on the locale. For example, `CST` could be U.S. `Central Standard Time` and `China Standard Time`. Such formats have been already deprecated in JDK, see [Three-letter time zone IDs](https://docs.oracle.com/javase/8/docs/api/java/util/TimeZone.html).
### Does this PR introduce any user-facing change?
No
### How was this patch tested?
By running `./dev/scalastyle`, and manual testing.
Closes#28051 from MaxGekk/doc-time-zone-option.
Authored-by: Maxim Gekk <max.gekk@gmail.com>
Signed-off-by: Wenchen Fan <wenchen@databricks.com>
### What changes were proposed in this pull request?
Based on the discussion in the mailing list [[Proposal] Modification to Spark's Semantic Versioning Policy](http://apache-spark-developers-list.1001551.n3.nabble.com/Proposal-Modification-to-Spark-s-Semantic-Versioning-Policy-td28938.html) , this PR is to add back the following APIs whose maintenance cost are relatively small.
- functions.toDegrees/toRadians
- functions.approxCountDistinct
- functions.monotonicallyIncreasingId
- Column.!==
- Dataset.explode
- Dataset.registerTempTable
- SQLContext.getOrCreate, setActive, clearActive, constructors
Below is the other removed APIs in the original PR, but not added back in this PR [https://issues.apache.org/jira/browse/SPARK-25908]:
- Remove some AccumulableInfo .apply() methods
- Remove non-label-specific multiclass precision/recall/fScore in favor of accuracy
- Remove unused Python StorageLevel constants
- Remove unused multiclass option in libsvm parsing
- Remove references to deprecated spark configs like spark.yarn.am.port
- Remove TaskContext.isRunningLocally
- Remove ShuffleMetrics.shuffle* methods
- Remove BaseReadWrite.context in favor of session
### Why are the changes needed?
Avoid breaking the APIs that are commonly used.
### Does this PR introduce any user-facing change?
Adding back the APIs that were removed in 3.0 branch does not introduce the user-facing changes, because Spark 3.0 has not been released.
### How was this patch tested?
Added a new test suite for these APIs.
Author: gatorsmile <gatorsmile@gmail.com>
Author: yi.wu <yi.wu@databricks.com>
Closes#27821 from gatorsmile/addAPIBackV2.
### What changes were proposed in this pull request?
This PR proposes to make pandas function APIs (`groupby.(cogroup.)applyInPandas` and `mapInPandas`) to ignore Python type hints.
### Why are the changes needed?
Python type hints are optional. It shouldn't affect where pandas UDFs are not used.
This is also a future work for them to support other type hints. We shouldn't at least throw an exception at this moment.
### Does this PR introduce any user-facing change?
No, it's master-only change.
```python
import pandas as pd
def pandas_plus_one(pdf: pd.DataFrame) -> pd.DataFrame:
return pdf + 1
spark.range(10).groupby('id').applyInPandas(pandas_plus_one, schema="id long").show()
```
```python
import pandas as pd
def pandas_plus_one(left: pd.DataFrame, right: pd.DataFrame) -> pd.DataFrame:
return left + 1
spark.range(10).groupby('id').cogroup(spark.range(10).groupby("id")).applyInPandas(pandas_plus_one, schema="id long").show()
```
```python
from typing import Iterator
import pandas as pd
def pandas_plus_one(iter: Iterator[pd.DataFrame]) -> Iterator[pd.DataFrame]:
return map(lambda v: v + 1, iter)
spark.range(10).mapInPandas(pandas_plus_one, schema="id long").show()
```
**Before:**
Exception
**After:**
```
+---+
| id|
+---+
| 1|
| 2|
| 3|
| 4|
| 5|
| 6|
| 7|
| 8|
| 9|
| 10|
+---+
```
### How was this patch tested?
Closes#28052 from HyukjinKwon/SPARK-31287.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
Based on the discussion in the mailing list [[Proposal] Modification to Spark's Semantic Versioning Policy](http://apache-spark-developers-list.1001551.n3.nabble.com/Proposal-Modification-to-Spark-s-Semantic-Versioning-Policy-td28938.html) , this PR is to add back the following APIs whose maintenance cost are relatively small.
- HiveContext
- createExternalTable APIs
### Why are the changes needed?
Avoid breaking the APIs that are commonly used.
### Does this PR introduce any user-facing change?
Adding back the APIs that were removed in 3.0 branch does not introduce the user-facing changes, because Spark 3.0 has not been released.
### How was this patch tested?
add a new test suite for createExternalTable APIs.
Closes#27815 from gatorsmile/addAPIsBack.
Lead-authored-by: gatorsmile <gatorsmile@gmail.com>
Co-authored-by: yi.wu <yi.wu@databricks.com>
Signed-off-by: gatorsmile <gatorsmile@gmail.com>
### What changes were proposed in this pull request?
Add ANOVATest and FValueTest to PySpark
### Why are the changes needed?
Parity between Scala and Python.
### Does this PR introduce any user-facing change?
Yes. Python ANOVATest and FValueTest
### How was this patch tested?
doctest
Closes#28012 from huaxingao/stats-python.
Authored-by: Huaxin Gao <huaxing@us.ibm.com>
Signed-off-by: zhengruifeng <ruifengz@foxmail.com>
### What changes were proposed in this pull request?
When `toPandas` API works on duplicate column names produced from operators like join, we see the error like:
```
ValueError: The truth value of a Series is ambiguous. Use a.empty, a.bool(), a.item(), a.any() or a.all().
```
This patch fixes the error in `toPandas` API.
### Why are the changes needed?
To make `toPandas` work on dataframe with duplicate column names.
### Does this PR introduce any user-facing change?
Yes. Previously calling `toPandas` API on a dataframe with duplicate column names will fail. After this patch, it will produce correct result.
### How was this patch tested?
Unit test.
Closes#28025 from viirya/SPARK-31186.
Authored-by: Liang-Chi Hsieh <viirya@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
the link for `partition discovery` is malformed, because for releases, there will contains` /docs/<version>/` in the full URL.
### Why are the changes needed?
fix doc
### Does this PR introduce any user-facing change?
no
### How was this patch tested?
`SKIP_SCALADOC=1 SKIP_RDOC=1 SKIP_SQLDOC=1 jekyll serve` locally verified
Closes#28017 from yaooqinn/doc.
Authored-by: Kent Yao <yaooqinn@hotmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
Fix errors and missing parts for datetime pattern document
1. The pattern we use is similar to DateTimeFormatter and SimpleDateFormat but not identical. So we shouldn't use any of them in the API docs but use a link to the doc of our own.
2. Some pattern letters are missing
3. Some pattern letters are explicitly banned - Set('A', 'c', 'e', 'n', 'N')
4. the second fraction pattern different logic for parsing and formatting
### Why are the changes needed?
fix and improve doc
### Does this PR introduce any user-facing change?
yes, new and updated doc
### How was this patch tested?
pass Jenkins
viewed locally with `jekyll serve`
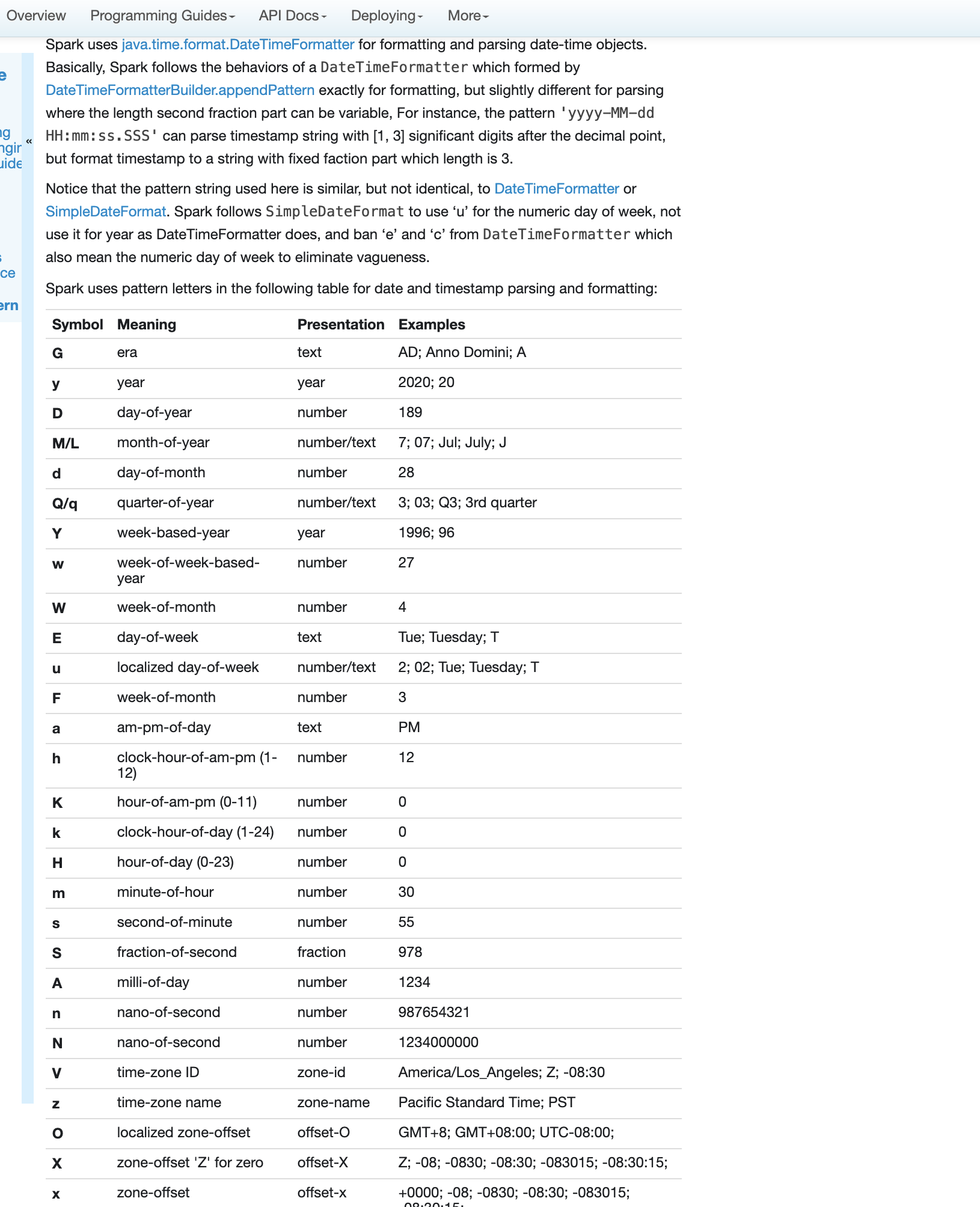
Closes#27956 from yaooqinn/SPARK-31189.
Authored-by: Kent Yao <yaooqinn@hotmail.com>
Signed-off-by: Wenchen Fan <wenchen@databricks.com>
### What changes were proposed in this pull request?
- Adds following overloaded variants to Scala `o.a.s.sql.functions`:
- `percentile_approx(e: Column, percentage: Array[Double], accuracy: Long): Column`
- `percentile_approx(columnName: String, percentage: Array[Double], accuracy: Long): Column`
- `percentile_approx(e: Column, percentage: Double, accuracy: Long): Column`
- `percentile_approx(columnName: String, percentage: Double, accuracy: Long): Column`
- `percentile_approx(e: Column, percentage: Seq[Double], accuracy: Long): Column` (primarily for
Python interop).
- `percentile_approx(columnName: String, percentage: Seq[Double], accuracy: Long): Column`
- Adds `percentile_approx` to `pyspark.sql.functions`.
- Adds `percentile_approx` function to SparkR.
### Why are the changes needed?
Currently we support `percentile_approx` only in SQL expression. It is inconvenient and makes this function relatively unknown.
### Does this PR introduce any user-facing change?
No.
### How was this patch tested?
New unit tests for SparkR an PySpark.
As for now there are no additional tests in Scala API ‒ `ApproximatePercentile` is well tested and Python (including docstrings) and R tests provide additional tests, so it seems unnecessary.
Closes#27278 from zero323/SPARK-30569.
Lead-authored-by: zero323 <mszymkiewicz@gmail.com>
Co-authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
jira link: https://issues.apache.org/jira/browse/SPARK-30930
Remove ML/MLLIB DeveloperApi annotations.
### Why are the changes needed?
The Developer APIs in ML/MLLIB have been there for a long time. They are stable now and are very unlikely to be changed or removed, so I unmark these Developer APIs in this PR.
### Does this PR introduce any user-facing change?
Yes. DeveloperApi annotations are removed from docs.
### How was this patch tested?
existing tests
Closes#27859 from huaxingao/spark-30930.
Authored-by: Huaxin Gao <huaxing@us.ibm.com>
Signed-off-by: Sean Owen <srowen@gmail.com>
### What changes were proposed in this pull request?
In Spark version 2.4 and earlier, datetime parsing, formatting and conversion are performed by using the hybrid calendar (Julian + Gregorian).
Since the Proleptic Gregorian calendar is de-facto calendar worldwide, as well as the chosen one in ANSI SQL standard, Spark 3.0 switches to it by using Java 8 API classes (the java.time packages that are based on ISO chronology ). The switching job is completed in SPARK-26651.
But after the switching, there are some patterns not compatible between Java 8 and Java 7, Spark needs its own definition on the patterns rather than depends on Java API.
In this PR, we achieve this by writing the document and shadow the incompatible letters. See more details in [SPARK-31030](https://issues.apache.org/jira/browse/SPARK-31030)
### Why are the changes needed?
For backward compatibility.
### Does this PR introduce any user-facing change?
No.
After we define our own datetime parsing and formatting patterns, it's same to old Spark version.
### How was this patch tested?
Existing and new added UT.
Locally document test:
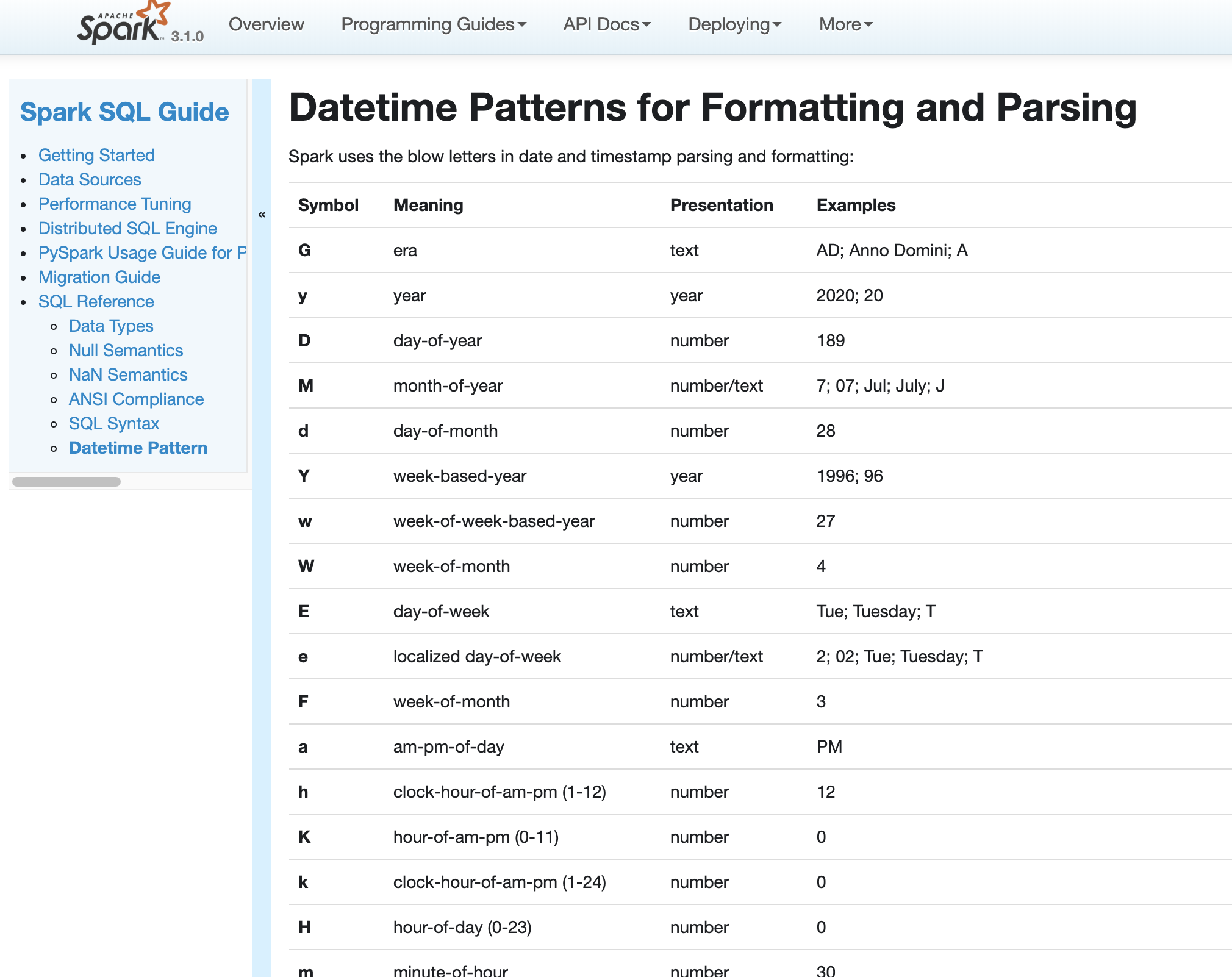
Closes#27830 from xuanyuanking/SPARK-31030.
Authored-by: Yuanjian Li <xyliyuanjian@gmail.com>
Signed-off-by: Wenchen Fan <wenchen@databricks.com>
### What changes were proposed in this pull request?
Adding a note to document `Row.asDict` behavior when there are duplicate fields.
### Why are the changes needed?
When a row contains duplicate fields, `asDict` and `_get_item_` behaves differently. We should document it to let users know the difference explicitly.
### Does this PR introduce any user-facing change?
No. Only document change.
### How was this patch tested?
Existing test.
Closes#27853 from viirya/SPARK-30941.
Authored-by: Liang-Chi Hsieh <viirya@gmail.com>
Signed-off-by: Dongjoon Hyun <dongjoon@apache.org>
### What changes were proposed in this pull request?
Updating ML docs for 3.0 changes
### Why are the changes needed?
I am auditing 3.0 ML changes, found some docs are missing or not updated. Need to update these.
### Does this PR introduce any user-facing change?
Yes, doc changes
### How was this patch tested?
Manually build and check
Closes#27762 from huaxingao/spark-doc.
Authored-by: Huaxin Gao <huaxing@us.ibm.com>
Signed-off-by: Sean Owen <srowen@gmail.com>
### What changes were proposed in this pull request?
Implement common base ML classes (`Predictor`, `PredictionModel`, `Classifier`, `ClasssificationModel` `ProbabilisticClassifier`, `ProbabilisticClasssificationModel`, `Regressor`, `RegrssionModel`) for non-Java backends.
Note
- `Predictor` and `JavaClassifier` should be abstract as `_fit` method is not implemented.
- `PredictionModel` should be abstract as `_transform` is not implemented.
### Why are the changes needed?
To provide extensions points for non-JVM algorithms, as well as a public (as opposed to `Java*` variants, which are commonly described in docstrings as private) hierarchy which can be used to distinguish between different classes of predictors.
For longer discussion see [SPARK-29212](https://issues.apache.org/jira/browse/SPARK-29212) and / or https://github.com/apache/spark/pull/25776.
### Does this PR introduce any user-facing change?
It adds new base classes as listed above, but effective interfaces (method resolution order notwithstanding) stay the same.
Additionally "private" `Java*` classes in`ml.regression` and `ml.classification` have been renamed to follow PEP-8 conventions (added leading underscore).
It is for discussion if the same should be done to equivalent classes from `ml.wrapper`.
If we take `JavaClassifier` as an example, type hierarchy will change from

to

Similarly the old model

will become

### How was this patch tested?
Existing unit tests.
Closes#27245 from zero323/SPARK-29212.
Authored-by: zero323 <mszymkiewicz@gmail.com>
Signed-off-by: zhengruifeng <ruifengz@foxmail.com>
### What changes were proposed in this pull request?
Current impl needs to convert ml.Vector to breeze.Vector, which can be skipped.
### Why are the changes needed?
avoid unnecessary vector conversions
### Does this PR introduce any user-facing change?
No
### How was this patch tested?
existing testsuites
Closes#27519 from zhengruifeng/gmm_transform_opt.
Authored-by: zhengruifeng <ruifengz@foxmail.com>
Signed-off-by: zhengruifeng <ruifengz@foxmail.com>
### What changes were proposed in this pull request?
Remove automatically resource coordination support from Standalone.
### Why are the changes needed?
Resource coordination is mainly designed for the scenario where multiple workers launched on the same host. However, it's, actually, a non-existed scenario for today's Spark. Because, Spark now can start multiple executors in a single Worker, while it only allow one executor per Worker at very beginning. So, now, it really help nothing for user to launch multiple workers on the same host. Thus, it's not worth for us to bring over complicated implementation and potential high maintain cost for such an impossible scenario.
### Does this PR introduce any user-facing change?
No, it's Spark 3.0 feature.
### How was this patch tested?
Pass Jenkins.
Closes#27722 from Ngone51/abandon_coordination.
Authored-by: yi.wu <yi.wu@databricks.com>
Signed-off-by: Xingbo Jiang <xingbo.jiang@databricks.com>
### What changes were proposed in this pull request?
This PR add Python API for invoking following higher functions:
- `transform`
- `exists`
- `forall`
- `filter`
- `aggregate`
- `zip_with`
- `transform_keys`
- `transform_values`
- `map_filter`
- `map_zip_with`
to `pyspark.sql`. Each of these accepts plain Python functions of one of the following types
- `(Column) -> Column: ...`
- `(Column, Column) -> Column: ...`
- `(Column, Column, Column) -> Column: ...`
Internally this proposal piggbacks on objects supporting Scala implementation ([SPARK-27297](https://issues.apache.org/jira/browse/SPARK-27297)) by:
1. Creating required `UnresolvedNamedLambdaVariables` exposing these as PySpark `Columns`
2. Invoking Python function with these columns as arguments.
3. Using the result, and underlying JVM objects from 1., to create `expressions.LambdaFunction` which is passed to desired expression, and repacked as Python `Column`.
### Why are the changes needed?
Currently higher order functions are available only using SQL and Scala API and can use only SQL expressions
```python
df.selectExpr("transform(values, x -> x + 1)")
```
This works reasonably well for simple functions, but can get really ugly with complex functions (complex functions, casts), resulting objects are somewhat verbose and we don't get any IDE support. Additionally DSL used, though very simple, is not documented.
With changes propose here, above query could be rewritten as:
```python
df.select(transform("values", lambda x: x + 1))
```
### Does this PR introduce any user-facing change?
No.
### How was this patch tested?
- For positive cases this PR adds doctest strings covering possible usage patterns.
- For negative cases (unsupported function types) this PR adds unit tests.
### Notes
If approved, the same approach can be used in SparkR.
Closes#27406 from zero323/SPARK-30681.
Authored-by: zero323 <mszymkiewicz@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This patch is to bump the master branch version to 3.1.0-SNAPSHOT.
### Why are the changes needed?
N/A
### Does this PR introduce any user-facing change?
N/A
### How was this patch tested?
N/A
Closes#27698 from gatorsmile/updateVersion.
Authored-by: gatorsmile <gatorsmile@gmail.com>
Signed-off-by: Dongjoon Hyun <dhyun@apple.com>
Fix for #27395
### What changes were proposed in this pull request?
The `allGather` method is added to the `BarrierTaskContext`. This method contains the same functionality as the `BarrierTaskContext.barrier` method; it blocks the task until all tasks make the call, at which time they may continue execution. In addition, the `allGather` method takes an input message. Upon returning from the `allGather` the task receives a list of all the messages sent by all the tasks that made the `allGather` call.
### Why are the changes needed?
There are many situations where having the tasks communicate in a synchronized way is useful. One simple example is if each task needs to start a server to serve requests from one another; first the tasks must find a free port (the result of which is undetermined beforehand) and then start making requests, but to do so they each must know the port chosen by the other task. An `allGather` method would allow them to inform each other of the port they will run on.
### Does this PR introduce any user-facing change?
Yes, an `BarrierTaskContext.allGather` method will be available through the Scala, Java, and Python APIs.
### How was this patch tested?
Most of the code path is already covered by tests to the `barrier` method, since this PR includes a refactor so that much code is shared by the `barrier` and `allGather` methods. However, a test is added to assert that an all gather on each tasks partition ID will return a list of every partition ID.
An example through the Python API:
```python
>>> from pyspark import BarrierTaskContext
>>>
>>> def f(iterator):
... context = BarrierTaskContext.get()
... return [context.allGather('{}'.format(context.partitionId()))]
...
>>> sc.parallelize(range(4), 4).barrier().mapPartitions(f).collect()[0]
[u'3', u'1', u'0', u'2']
```
Closes#27640 from sarthfrey/master.
Lead-authored-by: sarthfrey-db <sarth.frey@databricks.com>
Co-authored-by: sarthfrey <sarth.frey@gmail.com>
Signed-off-by: Xingbo Jiang <xingbo.jiang@databricks.com>
### What changes were proposed in this pull request?
The style of `EXPLAIN FORMATTED` output needs to be improved. We’ve already got some observations/ideas in
https://github.com/apache/spark/pull/27368#discussion_r376694496https://github.com/apache/spark/pull/27368#discussion_r376927143
Observations/Ideas:
1. Using comma as the separator is not clear, especially commas are used inside the expressions too.
2. Show the column counts first? For example, `Results [4]: …`
3. Currently the attribute names are automatically generated, this need to refined.
4. Add arguments field in common implementations as `EXPLAIN EXTENDED` did by calling `argString` in `TreeNode.simpleString`. This will eliminate most existing minor differences between
`EXPLAIN EXTENDED` and `EXPLAIN FORMATTED`.
5. Another improvement we can do is: the generated alias shouldn't include attribute id. collect_set(val, 0, 0)#123 looks clearer than collect_set(val#456, 0, 0)#123
This PR is currently addressing comments 2 & 4, and open for more discussions on improving readability.
### Why are the changes needed?
The readability of `EXPLAIN FORMATTED` need to be improved, which will help user better understand the query plan.
### Does this PR introduce any user-facing change?
Yes, `EXPLAIN FORMATTED` output style changed.
### How was this patch tested?
Update expect results of test cases in explain.sql
Closes#27509 from Eric5553/ExplainFormattedRefine.
Authored-by: Eric Wu <492960551@qq.com>
Signed-off-by: Wenchen Fan <wenchen@databricks.com>
### What changes were proposed in this pull request?
As discussed on the Jira ticket, this change clears the SQLContext._instantiatedContext class attribute when the SparkSession is stopped. That way, the attribute will be reset with a new, usable SQLContext when a new SparkSession is started.
### Why are the changes needed?
When the underlying SQLContext is instantiated for a SparkSession, the instance is saved as a class attribute and returned from subsequent calls to SQLContext.getOrCreate(). If the SparkContext is stopped and a new one started, the SQLContext class attribute is never cleared so any code which calls SQLContext.getOrCreate() will get a SQLContext with a reference to the old, unusable SparkContext.
A similar issue was identified and fixed for SparkSession in [SPARK-19055](https://issues.apache.org/jira/browse/SPARK-19055), but the fix did not change SQLContext as well. I ran into this because mllib still [uses](https://github.com/apache/spark/blob/master/python/pyspark/mllib/common.py#L105) SQLContext.getOrCreate() under the hood.
### Does this PR introduce any user-facing change?
No
### How was this patch tested?
A new test was added. I verified that the test fails without the included change.
Closes#27610 from afavaro/restart-sqlcontext.
Authored-by: Alex Favaro <alex.favaro@affirm.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
The `allGather` method is added to the `BarrierTaskContext`. This method contains the same functionality as the `BarrierTaskContext.barrier` method; it blocks the task until all tasks make the call, at which time they may continue execution. In addition, the `allGather` method takes an input message. Upon returning from the `allGather` the task receives a list of all the messages sent by all the tasks that made the `allGather` call.
### Why are the changes needed?
There are many situations where having the tasks communicate in a synchronized way is useful. One simple example is if each task needs to start a server to serve requests from one another; first the tasks must find a free port (the result of which is undetermined beforehand) and then start making requests, but to do so they each must know the port chosen by the other task. An `allGather` method would allow them to inform each other of the port they will run on.
### Does this PR introduce any user-facing change?
Yes, an `BarrierTaskContext.allGather` method will be available through the Scala, Java, and Python APIs.
### How was this patch tested?
Most of the code path is already covered by tests to the `barrier` method, since this PR includes a refactor so that much code is shared by the `barrier` and `allGather` methods. However, a test is added to assert that an all gather on each tasks partition ID will return a list of every partition ID.
An example through the Python API:
```python
>>> from pyspark import BarrierTaskContext
>>>
>>> def f(iterator):
... context = BarrierTaskContext.get()
... return [context.allGather('{}'.format(context.partitionId()))]
...
>>> sc.parallelize(range(4), 4).barrier().mapPartitions(f).collect()[0]
[u'3', u'1', u'0', u'2']
```
Closes#27395 from sarthfrey/master.
Lead-authored-by: sarthfrey-db <sarth.frey@databricks.com>
Co-authored-by: sarthfrey <sarth.frey@gmail.com>
Signed-off-by: Xiangrui Meng <meng@databricks.com>
(cherry picked from commit 57254c9719)
Signed-off-by: Xiangrui Meng <meng@databricks.com>
### What changes were proposed in this pull request?
This PR proposes to deprecate the APIs at `SQLContext` removed in SPARK-25908. We should remove equivalent APIs; however, seems we missed to deprecate.
While I am here, I fix one more issue. After SPARK-25908, `sc._jvm.SQLContext.getOrCreate` dose not exist anymore. So,
```python
from pyspark.sql import SQLContext
from pyspark import SparkContext
sc = SparkContext.getOrCreate()
SQLContext.getOrCreate(sc).range(10).show()
```
throws an exception as below:
```
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "/.../spark/python/pyspark/sql/context.py", line 110, in getOrCreate
jsqlContext = sc._jvm.SQLContext.getOrCreate(sc._jsc.sc())
File "/.../spark/python/lib/py4j-0.10.8.1-src.zip/py4j/java_gateway.py", line 1516, in __getattr__
py4j.protocol.Py4JError: org.apache.spark.sql.SQLContext.getOrCreate does not exist in the JVM
```
After this PR:
```
/.../spark/python/pyspark/sql/context.py:113: DeprecationWarning: Deprecated in 3.0.0. Use SparkSession.builder.getOrCreate() instead.
DeprecationWarning)
+---+
| id|
+---+
| 0|
| 1|
| 2|
| 3|
| 4|
| 5|
| 6|
| 7|
| 8|
| 9|
+---+
```
In case of the constructor of `SQLContext`, after this PR:
```python
from pyspark.sql import SQLContext
sc = SparkContext.getOrCreate()
SQLContext(sc)
```
```
/.../spark/python/pyspark/sql/context.py:77: DeprecationWarning: Deprecated in 3.0.0. Use SparkSession.builder.getOrCreate() instead.
DeprecationWarning)
```
### Why are the changes needed?
To promote to use SparkSession, and keep the API party consistent with Scala side.
### Does this PR introduce any user-facing change?
Yes, it will show deprecation warning to users.
### How was this patch tested?
Manually tested as described above. Unittests were also added.
Closes#27614 from HyukjinKwon/SPARK-30861.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
This commit is published into the public domain.
### What changes were proposed in this pull request?
Some syntax issues in docstrings have been fixed.
### Why are the changes needed?
In some places, the documentation did not render as intended, e.g. parameter documentations were not formatted as such.
### Does this PR introduce any user-facing change?
Slight improvements in documentation.
### How was this patch tested?
Manual testing. No new Sphinx warnings arise due to this change.
Closes#27613 from DavidToneian/SPARK-30859.
Authored-by: David Toneian <david@toneian.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR added two DeveloperApis to the Dataset[T] class. Both methods are just exposing lower-level methods to the Dataset[T] class.
### Why are the changes needed?
They are useful for checking whether two dataframes are the same when implementing dataframe caching in python, and also get a unique ID. It's easier to use if we wrap the lower-level APIs.
### Does this PR introduce any user-facing change?
```
scala> val df1 = Seq((1,2),(4,5)).toDF("col1", "col2")
df1: org.apache.spark.sql.DataFrame = [col1: int, col2: int]
scala> val df2 = Seq((1,2),(4,5)).toDF("col1", "col2")
df2: org.apache.spark.sql.DataFrame = [col1: int, col2: int]
scala> val df3 = Seq((0,2),(4,5)).toDF("col1", "col2")
df3: org.apache.spark.sql.DataFrame = [col1: int, col2: int]
scala> val df4 = Seq((0,2),(4,5)).toDF("col0", "col2")
df4: org.apache.spark.sql.DataFrame = [col0: int, col2: int]
scala> df1.semanticHash
res0: Int = 594427822
scala> df2.semanticHash
res1: Int = 594427822
scala> df1.sameSemantics(df2)
res2: Boolean = true
scala> df1.sameSemantics(df3)
res3: Boolean = false
scala> df3.semanticHash
res4: Int = -1592702048
scala> df4.semanticHash
res5: Int = -1592702048
scala> df4.sameSemantics(df3)
res6: Boolean = true
```
### How was this patch tested?
Unit test in scala and doctest in python.
Note: comments are copied from the corresponding lower-level APIs.
Note: There are some issues to be fixed that would improve the hash collision rate: https://github.com/apache/spark/pull/27565#discussion_r379881028Closes#27565 from liangz1/df-same-result.
Authored-by: Liang Zhang <liang.zhang@databricks.com>
Signed-off-by: WeichenXu <weichen.xu@databricks.com>
### What changes were proposed in this pull request?
This is a follow-up for #23124, add a new config `spark.sql.legacy.allowDuplicatedMapKeys` to control the behavior of removing duplicated map keys in build-in functions. With the default value `false`, Spark will throw a RuntimeException while duplicated keys are found.
### Why are the changes needed?
Prevent silent behavior changes.
### Does this PR introduce any user-facing change?
Yes, new config added and the default behavior for duplicated map keys changed to RuntimeException thrown.
### How was this patch tested?
Modify existing UT.
Closes#27478 from xuanyuanking/SPARK-25892-follow.
Authored-by: Yuanjian Li <xyliyuanjian@gmail.com>
Signed-off-by: Wenchen Fan <wenchen@databricks.com>
This commit is published into the public domain.
### What changes were proposed in this pull request?
This PR fixes the documentation of `DataFrameReader`, `DataFrameWriter`, `DataStreamReader`, and `DataStreamWriter`, where attributes of other classes were misrepresented as functions. Additionally, creation of hyperlinks across modules was fixed in these instances.
### Why are the changes needed?
The old state produced documentation that suggested invalid usage of PySpark objects (accessing attributes as though they were callable.)
### Does this PR introduce any user-facing change?
No, except for improved documentation.
### How was this patch tested?
No test added; documentation build runs through.
Closes#27553 from DavidToneian/docfix-DataFrameReader-DataFrameWriter-DataStreamReader-DataStreamWriter.
Authored-by: David Toneian <david@toneian.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
The `allGather` method is added to the `BarrierTaskContext`. This method contains the same functionality as the `BarrierTaskContext.barrier` method; it blocks the task until all tasks make the call, at which time they may continue execution. In addition, the `allGather` method takes an input message. Upon returning from the `allGather` the task receives a list of all the messages sent by all the tasks that made the `allGather` call.
### Why are the changes needed?
There are many situations where having the tasks communicate in a synchronized way is useful. One simple example is if each task needs to start a server to serve requests from one another; first the tasks must find a free port (the result of which is undetermined beforehand) and then start making requests, but to do so they each must know the port chosen by the other task. An `allGather` method would allow them to inform each other of the port they will run on.
### Does this PR introduce any user-facing change?
Yes, an `BarrierTaskContext.allGather` method will be available through the Scala, Java, and Python APIs.
### How was this patch tested?
Most of the code path is already covered by tests to the `barrier` method, since this PR includes a refactor so that much code is shared by the `barrier` and `allGather` methods. However, a test is added to assert that an all gather on each tasks partition ID will return a list of every partition ID.
An example through the Python API:
```python
>>> from pyspark import BarrierTaskContext
>>>
>>> def f(iterator):
... context = BarrierTaskContext.get()
... return [context.allGather('{}'.format(context.partitionId()))]
...
>>> sc.parallelize(range(4), 4).barrier().mapPartitions(f).collect()[0]
[u'3', u'1', u'0', u'2']
```
Closes#27395 from sarthfrey/master.
Lead-authored-by: sarthfrey-db <sarth.frey@databricks.com>
Co-authored-by: sarthfrey <sarth.frey@gmail.com>
Signed-off-by: Xiangrui Meng <meng@databricks.com>
### What changes were proposed in this pull request?
In this PR, we add a parameter in the python function vector_to_array(col) that allows converting to a column of arrays of Float (32bits) in scala, which would be mapped to a numpy array of dtype=float32.
### Why are the changes needed?
In the downstream ML training, using float32 instead of float64 (default) would allow a larger batch size, i.e., allow more data to fit in the memory.
### Does this PR introduce any user-facing change?
Yes.
Old: `vector_to_array()` only take one param
```
df.select(vector_to_array("colA"), ...)
```
New: `vector_to_array()` can take an additional optional param: `dtype` = "float32" (or "float64")
```
df.select(vector_to_array("colA", "float32"), ...)
```
### How was this patch tested?
Unit test in scala.
doctest in python.
Closes#27522 from liangz1/udf-float32.
Authored-by: Liang Zhang <liang.zhang@databricks.com>
Signed-off-by: WeichenXu <weichen.xu@databricks.com>
### What changes were proposed in this pull request?
This is another PR for stage level scheduling. In particular this adds changes to the dynamic allocation manager and the scheduler backend to be able to track what executors are needed per ResourceProfile. Note the api is still private to Spark until the entire feature gets in, so this functionality will be there but only usable by tests for profiles other then the DefaultProfile.
The main changes here are simply tracking things on a ResourceProfile basis as well as sending the executor requests to the scheduler backend for all ResourceProfiles.
I introduce a ResourceProfileManager in this PR that will track all the actual ResourceProfile objects so that we can keep them all in a single place and just pass around and use in datastructures the resource profile id. The resource profile id can be used with the ResourceProfileManager to get the actual ResourceProfile contents.
There are various places in the code that use executor "slots" for things. The ResourceProfile adds functionality to keep that calculation in it. This logic is more complex then it should due to standalone mode and mesos coarse grained not setting the executor cores config. They default to all cores on the worker, so calculating slots is harder there.
This PR keeps the functionality to make the cores the limiting resource because the scheduler still uses that for "slots" for a few things.
This PR does also add the resource profile id to the Stage and stage info classes to be able to test things easier. That full set of changes will come with the scheduler PR that will be after this one.
The PR stops at the scheduler backend pieces for the cluster manager and the real YARN support hasn't been added in this PR, that again will be in a separate PR, so this has a few of the API changes up to the cluster manager and then just uses the default profile requests to continue.
The code for the entire feature is here for reference: https://github.com/apache/spark/pull/27053/files although it needs to be upmerged again as well.
### Why are the changes needed?
Needed for stage level scheduling feature.
### Does this PR introduce any user-facing change?
No user facing api changes added here.
### How was this patch tested?
Lots of unit tests and manually testing. I tested on yarn, k8s, standalone, local modes. Ran both failure and success cases.
Closes#27313 from tgravescs/SPARK-29148.
Authored-by: Thomas Graves <tgraves@nvidia.com>
Signed-off-by: Thomas Graves <tgraves@apache.org>
### What changes were proposed in this pull request?
This PR targets to document the Pandas UDF redesign with type hints introduced at SPARK-28264.
Mostly self-describing; however, there are few things to note for reviewers.
1. This PR replace the existing documentation of pandas UDFs to the newer redesign to promote the Python type hints. I added some words that Spark 3.0 still keeps the compatibility though.
2. This PR proposes to name non-pandas UDFs as "Pandas Function API"
3. SCALAR_ITER become two separate sections to reduce confusion:
- `Iterator[pd.Series]` -> `Iterator[pd.Series]`
- `Iterator[Tuple[pd.Series, ...]]` -> `Iterator[pd.Series]`
4. I removed some examples that look overkill to me.
5. I also removed some information in the doc, that seems duplicating or too much.
### Why are the changes needed?
To document new redesign in pandas UDF.
### Does this PR introduce any user-facing change?
No.
### How was this patch tested?
Existing tests should cover.
Closes#27466 from HyukjinKwon/SPARK-30722.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
Fix PySpark test failures for using Pandas >= 1.0.0.
### Why are the changes needed?
Pandas 1.0.0 has recently been released and has API changes that result in PySpark test failures, this PR fixes the broken tests.
### Does this PR introduce any user-facing change?
No
### How was this patch tested?
Manually tested with Pandas 1.0.1 and PyArrow 0.16.0
Closes#27529 from BryanCutler/pandas-fix-tests-1.0-SPARK-30777.
Authored-by: Bryan Cutler <cutlerb@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
Add ```HasBlockSize``` in shared Params in both Scala and Python.
Make ALS/MLP extend ```HasBlockSize```
### Why are the changes needed?
Add ```HasBlockSize ``` in ALS, so user can specify the blockSize.
Make ```HasBlockSize``` a shared param so both ALS and MLP can use it.
### Does this PR introduce any user-facing change?
Yes
```ALS.setBlockSize/getBlockSize```
```ALSModel.setBlockSize/getBlockSize```
### How was this patch tested?
Manually tested. Also added doctest.
Closes#27501 from huaxingao/spark_30662.
Authored-by: Huaxin Gao <huaxing@us.ibm.com>
Signed-off-by: zhengruifeng <ruifengz@foxmail.com>
### What changes were proposed in this pull request?
Revert
#27360#27396#27374#27389
### Why are the changes needed?
BLAS need more performace tests, specially on sparse datasets.
Perfermance test of LogisticRegression (https://github.com/apache/spark/pull/27374) on sparse dataset shows that blockify vectors to matrices and use BLAS will cause performance regression.
LinearSVC and LinearRegression were also updated in the same way as LogisticRegression, so we need to revert them to make sure no regression.
### Does this PR introduce any user-facing change?
remove newly added param blockSize
### How was this patch tested?
reverted testsuites
Closes#27487 from zhengruifeng/revert_blockify_ii.
Authored-by: zhengruifeng <ruifengz@foxmail.com>
Signed-off-by: zhengruifeng <ruifengz@foxmail.com>
### What changes were proposed in this pull request?
This PR fixes some typos in `python/pyspark/sql/types.py` file.
### Why are the changes needed?
To deliver correct wording in documentation and codes.
### Does this PR introduce any user-facing change?
Yes, it fixes some typos in user-facing API documentation.
### How was this patch tested?
Locally tested the linter.
Closes#27475 from sharifahmad2061/master.
Lead-authored-by: sharif ahmad <sharifahmad2061@gmail.com>
Co-authored-by: Sharif ahmad <sharifahmad2061@users.noreply.github.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR renames `spark.sql.pandas.udf.buffer.size` to `spark.sql.execution.pandas.udf.buffer.size` to be more consistent with other pandas configuration prefixes, given:
- `spark.sql.execution.pandas.arrowSafeTypeConversion`
- `spark.sql.execution.pandas.respectSessionTimeZone`
- `spark.sql.legacy.execution.pandas.groupedMap.assignColumnsByName`
- other configurations like `spark.sql.execution.arrow.*`.
### Why are the changes needed?
To make configuration names consistent.
### Does this PR introduce any user-facing change?
No because this configuration was not released yet.
### How was this patch tested?
Existing tests should cover.
Closes#27450 from HyukjinKwon/SPARK-27870-followup.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR aims to increase the timeout of `StreamingLogisticRegressionWithSGDTests.test_parameter_accuracy` from 30s (default) to 60s.
In this PR, before increasing the timeout,
1. I verified that this is not a JDK11 environmental issue by repeating 3 times first.
2. I reproduced the accuracy failure by reducing the timeout in Jenkins (https://github.com/apache/spark/pull/27424#issuecomment-580981262)
Then, the final commit passed the Jenkins.
### Why are the changes needed?
This seems to happen when Jenkins environment has congestion and the jobs are slowdown. The streaming job seems to be unable to repeat the designed iteration `numIteration=25` in 30 seconds. Since the error is decreasing at each iteration, the failure occurs.
By reducing the timeout, we can reproduce the similar issue locally like Jenkins.
```python
- eventually(condition, catch_assertions=True)
+ eventually(condition, timeout=10.0, catch_assertions=True)
```
```
$ python/run-tests --testname 'pyspark.mllib.tests.test_streaming_algorithms StreamingLogisticRegressionWithSGDTests.test_parameter_accuracy' --python-executables=python
...
======================================================================
FAIL: test_parameter_accuracy (pyspark.mllib.tests.test_streaming_algorithms.StreamingLogisticRegressionWithSGDTests)
----------------------------------------------------------------------
Traceback (most recent call last):
File "/Users/dongjoon/PRS/SPARK-TEST/python/pyspark/mllib/tests/test_streaming_algorithms.py", line 229, in test_parameter_accuracy
eventually(condition, timeout=10.0, catch_assertions=True)
File "/Users/dongjoon/PRS/SPARK-TEST/python/pyspark/testing/utils.py", line 86, in eventually
raise lastValue
Reproduce the error
File "/Users/dongjoon/PRS/SPARK-TEST/python/pyspark/testing/utils.py", line 77, in eventually
lastValue = condition()
File "/Users/dongjoon/PRS/SPARK-TEST/python/pyspark/mllib/tests/test_streaming_algorithms.py", line 226, in condition
self.assertAlmostEqual(rel, 0.1, 1)
AssertionError: 0.25749106949322637 != 0.1 within 1 places (0.15749106949322636 difference)
----------------------------------------------------------------------
Ran 1 test in 14.814s
FAILED (failures=1)
```
### Does this PR introduce any user-facing change?
No.
### How was this patch tested?
Pass the Jenkins (and manual check by reducing the timeout).
Since this is a flakiness issue depending on the Jenkins job situation, it's difficult to reproduce there.
Closes#27424 from dongjoon-hyun/SPARK-TEST.
Authored-by: Dongjoon Hyun <dhyun@apple.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
1, use blocks instead of vectors for performance improvement
2, use Level-2 BLAS
3, move standardization of input vectors outside of gradient computation
### Why are the changes needed?
1, less RAM to persist training data; (save ~40%)
2, faster than existing impl; (30% ~ 102%)
### Does this PR introduce any user-facing change?
add a new expert param `blockSize`
### How was this patch tested?
updated testsuites
Closes#27396 from zhengruifeng/blockify_lireg.
Authored-by: zhengruifeng <ruifengz@foxmail.com>
Signed-off-by: Sean Owen <srowen@gmail.com>
### What changes were proposed in this pull request?
remove duplicate setter in ```BucketedRandomProjectionLSH```
### Why are the changes needed?
Remove the duplicate ```setInputCol/setOutputCol``` in ```BucketedRandomProjectionLSH``` because these two setter are already in super class ```LSH```
### Does this PR introduce any user-facing change?
No
### How was this patch tested?
Manually checked.
Closes#27397 from huaxingao/spark-29093.
Authored-by: Huaxin Gao <huaxing@us.ibm.com>
Signed-off-by: Dongjoon Hyun <dhyun@apple.com>
### What changes were proposed in this pull request?
Make ALS/MLP extend ```HasBlockSize```
### Why are the changes needed?
Currently, MLP has its own ```blockSize``` param, we should make MLP extend ```HasBlockSize``` since ```HasBlockSize``` was added in ```sharedParams.scala``` recently.
ALS doesn't have ```blockSize``` param now, we can make it extend ```HasBlockSize```, so user can specify the ```blockSize```.
### Does this PR introduce any user-facing change?
Yes
```ALS.setBlockSize``` and ```ALS.getBlockSize```
```ALSModel.setBlockSize``` and ```ALSModel.getBlockSize```
### How was this patch tested?
Manually tested. Also added doctest.
Closes#27389 from huaxingao/spark-30662.
Authored-by: Huaxin Gao <huaxing@us.ibm.com>
Signed-off-by: Sean Owen <srowen@gmail.com>
### What changes were proposed in this pull request?
1, use blocks instead of vectors
2, use Level-2 BLAS for binary, use Level-3 BLAS for multinomial
### Why are the changes needed?
1, less RAM to persist training data; (save ~40%)
2, faster than existing impl; (40% ~ 92%)
### Does this PR introduce any user-facing change?
add a new expert param `blockSize`
### How was this patch tested?
updated testsuites
Closes#27374 from zhengruifeng/blockify_lor.
Authored-by: zhengruifeng <ruifengz@foxmail.com>
Signed-off-by: Sean Owen <srowen@gmail.com>
### What changes were proposed in this pull request?
1, stack input vectors to blocks (like ALS/MLP);
2, add new param `blockSize`;
3, add a new class `InstanceBlock`
4, standardize the input outside of optimization procedure;
### Why are the changes needed?
1, reduce RAM to persist traing dataset; (save ~40% in test)
2, use Level-2 BLAS routines; (12% ~ 28% faster, without native BLAS)
### Does this PR introduce any user-facing change?
a new param `blockSize`
### How was this patch tested?
existing and updated testsuites
Closes#27360 from zhengruifeng/blockify_svc.
Authored-by: zhengruifeng <ruifengz@foxmail.com>
Signed-off-by: zhengruifeng <ruifengz@foxmail.com>
### What changes were proposed in this pull request?
Prevent unnecessary copies of data during conversion from Arrow to Pandas.
### Why are the changes needed?
During conversion of pyarrow data to Pandas, columns are checked for timestamp types and then modified to correct for local timezone. If the data contains no timestamp types, then unnecessary copies of the data can be made. This is most prevalent when checking columns of a pandas DataFrame where each series is assigned back to the DataFrame, regardless if it had timestamps. See https://www.mail-archive.com/devarrow.apache.org/msg17008.html and ARROW-7596 for discussion.
### Does this PR introduce any user-facing change?
No
### How was this patch tested?
Existing tests
Closes#27358 from BryanCutler/pyspark-pandas-timestamp-copy-fix-SPARK-30640.
Authored-by: Bryan Cutler <cutlerb@gmail.com>
Signed-off-by: Bryan Cutler <cutlerb@gmail.com>
This reverts commit 1d20d13149.
Closes#27351 from gatorsmile/revertSPARK25496.
Authored-by: Xiao Li <gatorsmile@gmail.com>
Signed-off-by: Dongjoon Hyun <dhyun@apple.com>
### What changes were proposed in this pull request?
Minor documentation fix
### Why are the changes needed?
### Does this PR introduce any user-facing change?
### How was this patch tested?
Manually; consider adding tests?
Closes#27295 from deepyaman/patch-2.
Authored-by: Deepyaman Datta <deepyaman.datta@utexas.edu>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
add a param `bootstrap` to control whether bootstrap samples are used.
### Why are the changes needed?
Current RF with numTrees=1 will directly build a tree using the orignial dataset,
while with numTrees>1 it will use bootstrap samples to build trees.
This design is for training a DecisionTreeModel by the impl of RandomForest, however, it is somewhat strange.
In Scikit-Learn, there is a param [bootstrap](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html#sklearn.ensemble.RandomForestClassifier) to control whether bootstrap samples are used.
### Does this PR introduce any user-facing change?
Yes, new param is added
### How was this patch tested?
existing testsuites
Closes#27254 from zhengruifeng/add_bootstrap.
Authored-by: zhengruifeng <ruifengz@foxmail.com>
Signed-off-by: zhengruifeng <ruifengz@foxmail.com>
### What changes were proposed in this pull request?
This PR adds:
- `pyspark.sql.functions.overlay` function to PySpark
- `overlay` function to SparkR
### Why are the changes needed?
Feature parity. At the moment R and Python users can access this function only using SQL or `expr` / `selectExpr`.
### Does this PR introduce any user-facing change?
No.
### How was this patch tested?
New unit tests.
Closes#27325 from zero323/SPARK-30607.
Authored-by: zero323 <mszymkiewicz@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR proposes to redesign pandas UDFs as described in [the proposal](https://docs.google.com/document/d/1-kV0FS_LF2zvaRh_GhkV32Uqksm_Sq8SvnBBmRyxm30/edit?usp=sharing).
```python
from pyspark.sql.functions import pandas_udf
import pandas as pd
pandas_udf("long")
def plug_one(s: pd.Series) -> pd.Series:
return s + 1
spark.range(10).select(plug_one("id")).show()
```
```
+------------+
|plug_one(id)|
+------------+
| 1|
| 2|
| 3|
| 4|
| 5|
| 6|
| 7|
| 8|
| 9|
| 10|
+------------+
```
Note that, this PR address one of the future improvements described [here](https://docs.google.com/document/d/1-kV0FS_LF2zvaRh_GhkV32Uqksm_Sq8SvnBBmRyxm30/edit#heading=h.h3ncjpk6ujqu), "A couple of less-intuitive pandas UDF types" (by zero323) together.
In short,
- Adds new way with type hints as an alternative and experimental way.
```python
pandas_udf(schema='...')
def func(c1: Series, c2: Series) -> DataFrame:
pass
```
- Replace and/or add an alias for three types below from UDF, and make them as separate standalone APIs. So, `pandas_udf` is now consistent with regular `udf`s and other expressions.
`df.mapInPandas(udf)` -replace-> `df.mapInPandas(f, schema)`
`df.groupby.apply(udf)` -alias-> `df.groupby.applyInPandas(f, schema)`
`df.groupby.cogroup.apply(udf)` -replace-> `df.groupby.cogroup.applyInPandas(f, schema)`
*`df.groupby.apply` was added from 2.3 while the other were added in the master only.
- No deprecation for the existing ways for now.
```python
pandas_udf(schema='...', functionType=PandasUDFType.SCALAR)
def func(c1, c2):
pass
```
If users are happy with this, I plan to deprecate the existing way and declare using type hints is not experimental anymore.
One design goal in this PR was that, avoid touching the internal (since we didn't deprecate the old ways for now), but supports type hints with a minimised changes only at the interface.
- Once we deprecate or remove the old ways, I think it requires another refactoring for the internal in the future. At the very least, we should rename internal pandas evaluation types.
- If users find this experimental type hints isn't quite helpful, we should simply revert the changes at the interface level.
### Why are the changes needed?
In order to address old design issues. Please see [the proposal](https://docs.google.com/document/d/1-kV0FS_LF2zvaRh_GhkV32Uqksm_Sq8SvnBBmRyxm30/edit?usp=sharing).
### Does this PR introduce any user-facing change?
For behaviour changes, No.
It adds new ways to use pandas UDFs by using type hints. See below.
**SCALAR**:
```python
pandas_udf(schema='...')
def func(c1: Series, c2: DataFrame) -> Series:
pass # DataFrame represents a struct column
```
**SCALAR_ITER**:
```python
pandas_udf(schema='...')
def func(iter: Iterator[Tuple[Series, DataFrame, ...]]) -> Iterator[Series]:
pass # Same as SCALAR but wrapped by Iterator
```
**GROUPED_AGG**:
```python
pandas_udf(schema='...')
def func(c1: Series, c2: DataFrame) -> int:
pass # DataFrame represents a struct column
```
**GROUPED_MAP**:
This was added in Spark 2.3 as of SPARK-20396. As described above, it keeps the existing behaviour. Additionally, we now have a new alias `groupby.applyInPandas` for `groupby.apply`. See the example below:
```python
def func(pdf):
return pdf
df.groupby("...").applyInPandas(func, schema=df.schema)
```
**MAP_ITER**: this is not a pandas UDF anymore
This was added in Spark 3.0 as of SPARK-28198; and this PR replaces the usages. See the example below:
```python
def func(iter):
for df in iter:
yield df
df.mapInPandas(func, df.schema)
```
**COGROUPED_MAP**: this is not a pandas UDF anymore
This was added in Spark 3.0 as of SPARK-27463; and this PR replaces the usages. See the example below:
```python
def asof_join(left, right):
return pd.merge_asof(left, right, on="...", by="...")
df1.groupby("...").cogroup(df2.groupby("...")).applyInPandas(asof_join, schema="...")
```
### How was this patch tested?
Unittests added and tested against Python 2.7, 3.6 and 3.7.
Closes#27165 from HyukjinKwon/revisit-pandas.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR propose to disallow negative `scale` of `Decimal` in Spark. And this PR brings two behavior changes:
1) for literals like `1.23E4BD` or `1.23E4`(with `spark.sql.legacy.exponentLiteralAsDecimal.enabled`=true, see [SPARK-29956](https://issues.apache.org/jira/browse/SPARK-29956)), we set its `(precision, scale)` to (5, 0) rather than (3, -2);
2) add negative `scale` check inside the decimal method if it exposes to set `scale` explicitly. If check fails, `AnalysisException` throws.
And user could still use `spark.sql.legacy.allowNegativeScaleOfDecimal.enabled` to restore the previous behavior.
### Why are the changes needed?
According to SQL standard,
> 4.4.2 Characteristics of numbers
An exact numeric type has a precision P and a scale S. P is a positive integer that determines the number of significant digits in a particular radix R, where R is either 2 or 10. S is a non-negative integer.
scale of Decimal should always be non-negative. And other mainstream databases, like Presto, PostgreSQL, also don't allow negative scale.
Presto:
```
presto:default> create table t (i decimal(2, -1));
Query 20191213_081238_00017_i448h failed: line 1:30: mismatched input '-'. Expecting: <integer>, <type>
create table t (i decimal(2, -1))
```
PostgrelSQL:
```
postgres=# create table t(i decimal(2, -1));
ERROR: NUMERIC scale -1 must be between 0 and precision 2
LINE 1: create table t(i decimal(2, -1));
^
```
And, actually, Spark itself already doesn't allow to create table with negative decimal types using SQL:
```
scala> spark.sql("create table t(i decimal(2, -1))");
org.apache.spark.sql.catalyst.parser.ParseException:
no viable alternative at input 'create table t(i decimal(2, -'(line 1, pos 28)
== SQL ==
create table t(i decimal(2, -1))
----------------------------^^^
at org.apache.spark.sql.catalyst.parser.ParseException.withCommand(ParseDriver.scala:263)
at org.apache.spark.sql.catalyst.parser.AbstractSqlParser.parse(ParseDriver.scala:130)
at org.apache.spark.sql.execution.SparkSqlParser.parse(SparkSqlParser.scala:48)
at org.apache.spark.sql.catalyst.parser.AbstractSqlParser.parsePlan(ParseDriver.scala:76)
at org.apache.spark.sql.SparkSession.$anonfun$sql$1(SparkSession.scala:605)
at org.apache.spark.sql.catalyst.QueryPlanningTracker.measurePhase(QueryPlanningTracker.scala:111)
at org.apache.spark.sql.SparkSession.sql(SparkSession.scala:605)
... 35 elided
```
However, it is still possible to create such table or `DatFrame` using Spark SQL programming API:
```
scala> val tb =
CatalogTable(
TableIdentifier("test", None),
CatalogTableType.MANAGED,
CatalogStorageFormat.empty,
StructType(StructField("i", DecimalType(2, -1) ) :: Nil))
```
```
scala> spark.sql("SELECT 1.23E4BD")
res2: org.apache.spark.sql.DataFrame = [1.23E+4: decimal(3,-2)]
```
while, these two different behavior could make user confused.
On the other side, even if user creates such table or `DataFrame` with negative scale decimal type, it can't write data out if using format, like `parquet` or `orc`. Because these formats have their own check for negative scale and fail on it.
```
scala> spark.sql("SELECT 1.23E4BD").write.saveAsTable("parquet")
19/12/13 17:37:04 ERROR Executor: Exception in task 0.0 in stage 0.0 (TID 0)
java.lang.IllegalArgumentException: Invalid DECIMAL scale: -2
at org.apache.parquet.Preconditions.checkArgument(Preconditions.java:53)
at org.apache.parquet.schema.Types$BasePrimitiveBuilder.decimalMetadata(Types.java:495)
at org.apache.parquet.schema.Types$BasePrimitiveBuilder.build(Types.java:403)
at org.apache.parquet.schema.Types$BasePrimitiveBuilder.build(Types.java:309)
at org.apache.parquet.schema.Types$Builder.named(Types.java:290)
at org.apache.spark.sql.execution.datasources.parquet.SparkToParquetSchemaConverter.convertField(ParquetSchemaConverter.scala:428)
at org.apache.spark.sql.execution.datasources.parquet.SparkToParquetSchemaConverter.convertField(ParquetSchemaConverter.scala:334)
at org.apache.spark.sql.execution.datasources.parquet.SparkToParquetSchemaConverter.$anonfun$convert$2(ParquetSchemaConverter.scala:326)
at scala.collection.TraversableLike.$anonfun$map$1(TraversableLike.scala:238)
at scala.collection.Iterator.foreach(Iterator.scala:941)
at scala.collection.Iterator.foreach$(Iterator.scala:941)
at scala.collection.AbstractIterator.foreach(Iterator.scala:1429)
at scala.collection.IterableLike.foreach(IterableLike.scala:74)
at scala.collection.IterableLike.foreach$(IterableLike.scala:73)
at org.apache.spark.sql.types.StructType.foreach(StructType.scala:99)
at scala.collection.TraversableLike.map(TraversableLike.scala:238)
at scala.collection.TraversableLike.map$(TraversableLike.scala:231)
at org.apache.spark.sql.types.StructType.map(StructType.scala:99)
at org.apache.spark.sql.execution.datasources.parquet.SparkToParquetSchemaConverter.convert(ParquetSchemaConverter.scala:326)
at org.apache.spark.sql.execution.datasources.parquet.ParquetWriteSupport.init(ParquetWriteSupport.scala:97)
at org.apache.parquet.hadoop.ParquetOutputFormat.getRecordWriter(ParquetOutputFormat.java:388)
at org.apache.parquet.hadoop.ParquetOutputFormat.getRecordWriter(ParquetOutputFormat.java:349)
at org.apache.spark.sql.execution.datasources.parquet.ParquetOutputWriter.<init>(ParquetOutputWriter.scala:37)
at org.apache.spark.sql.execution.datasources.parquet.ParquetFileFormat$$anon$1.newInstance(ParquetFileFormat.scala:150)
at org.apache.spark.sql.execution.datasources.SingleDirectoryDataWriter.newOutputWriter(FileFormatDataWriter.scala:124)
at org.apache.spark.sql.execution.datasources.SingleDirectoryDataWriter.<init>(FileFormatDataWriter.scala:109)
at org.apache.spark.sql.execution.datasources.FileFormatWriter$.executeTask(FileFormatWriter.scala:264)
at org.apache.spark.sql.execution.datasources.FileFormatWriter$.$anonfun$write$15(FileFormatWriter.scala:205)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:127)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:441)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1377)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:444)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
at java.lang.Thread.run(Thread.java:748)
```
So, I think it would be better to disallow negative scale totally and make behaviors above be consistent.
### Does this PR introduce any user-facing change?
Yes, if `spark.sql.legacy.allowNegativeScaleOfDecimal.enabled=false`, user couldn't create Decimal value with negative scale anymore.
### How was this patch tested?
Added new tests in `ExpressionParserSuite` and `DecimalSuite`;
Updated `SQLQueryTestSuite`.
Closes#26881 from Ngone51/nonnegative-scale.
Authored-by: yi.wu <yi.wu@databricks.com>
Signed-off-by: Wenchen Fan <wenchen@databricks.com>
### What changes were proposed in this pull request?
https://github.com/apache/spark/pull/26809 added `Dataset.tail` API. It should be good to have it in PySpark API as well.
### Why are the changes needed?
To support consistent APIs.
### Does this PR introduce any user-facing change?
No. It adds a new API.
### How was this patch tested?
Manually tested and doctest was added.
Closes#27251 from HyukjinKwon/SPARK-30539.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: Dongjoon Hyun <dhyun@apple.com>
### What changes were proposed in this pull request?
This PR adds:
- `pyspark.ml.regression.JavaRegressor`
- `pyspark.ml.regression.JavaRegressionModel`
classes and replaces `JavaPredictor` and `JavaPredictionModel` in
- `LinearRegression` / `LinearRegressionModel`
- `DecisionTreeRegressor` / `DecisionTreeRegressionModel` (just addition as `JavaPredictionModel` hasn't been used)
- `RandomForestRegressor` / `RandomForestRegressionModel` (just addition as `JavaPredictionModel` hasn't been used)
- `GBTRegressor` / `GBTRegressionModel` (just addition as `JavaPredictionModel` hasn't been used)
- `AFTSurvivalRegression` / `AFTSurvivalRegressionModel`
- `GeneralizedLinearRegression` / `GeneralizedLinearRegressionModel`
- `FMRegressor` / `FMRegressionModel`
### Why are the changes needed?
- Internal PySpark consistency.
- Feature parity with Scala.
- Intermediate step towards implementing [SPARK-29212](https://issues.apache.org/jira/browse/SPARK-29212)
### Does this PR introduce any user-facing change?
It adds new base classes, so it will affect `mro`. Otherwise interfaces should stay intact.
### How was this patch tested?
Existing tests.
Closes#27241 from zero323/SPARK-30533.
Authored-by: zero323 <mszymkiewicz@gmail.com>
Signed-off-by: Sean Owen <srowen@gmail.com>
### What changes were proposed in this pull request?
Another followup of 4398dfa709
I missed two more tests added:
```
======================================================================
ERROR [0.133s]: test_to_pandas_from_mixed_dataframe (pyspark.sql.tests.test_dataframe.DataFrameTests)
----------------------------------------------------------------------
Traceback (most recent call last):
File "/home/jenkins/python/pyspark/sql/tests/test_dataframe.py", line 617, in test_to_pandas_from_mixed_dataframe
self.assertTrue(np.all(pdf_with_only_nulls.dtypes == pdf_with_some_nulls.dtypes))
AssertionError: False is not true
======================================================================
ERROR [0.061s]: test_to_pandas_from_null_dataframe (pyspark.sql.tests.test_dataframe.DataFrameTests)
----------------------------------------------------------------------
Traceback (most recent call last):
File "/home/jenkins/python/pyspark/sql/tests/test_dataframe.py", line 588, in test_to_pandas_from_null_dataframe
self.assertEqual(types[0], np.float64)
AssertionError: dtype('O') != <class 'numpy.float64'>
----------------------------------------------------------------------
```
### Why are the changes needed?
To make the test independent of default values of configuration.
### Does this PR introduce any user-facing change?
No.
### How was this patch tested?
Manually tested and Jenkins should test.
Closes#27250 from HyukjinKwon/SPARK-29188-followup2.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR proposes to explicitly disable Arrow execution for the test of toPandas empty types. If `spark.sql.execution.arrow.pyspark.enabled` is enabled by default, this test alone fails as below:
```
======================================================================
ERROR [0.205s]: test_to_pandas_from_empty_dataframe (pyspark.sql.tests.test_dataframe.DataFrameTests)
----------------------------------------------------------------------
Traceback (most recent call last):
File "/.../pyspark/sql/tests/test_dataframe.py", line 568, in test_to_pandas_from_empty_dataframe
self.assertTrue(np.all(dtypes_when_empty_df == dtypes_when_nonempty_df))
AssertionError: False is not true
----------------------------------------------------------------------
```
it should be best to explicitly disable for the test that only works when it's disabled.
### Why are the changes needed?
To make the test independent of default values of configuration.
### Does this PR introduce any user-facing change?
No.
### How was this patch tested?
Manually tested and Jenkins should test.
Closes#27247 from HyukjinKwon/SPARK-29188-followup.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: Dongjoon Hyun <dhyun@apple.com>
### What changes were proposed in this pull request?
In the PR, I propose to remove the SQL config `spark.sql.execution.pandas.respectSessionTimeZone` which has been deprecated since Spark 2.3.
### Why are the changes needed?
To improve code maintainability.
### Does this PR introduce any user-facing change?
Yes.
### How was this patch tested?
by running python tests, https://spark.apache.org/docs/latest/building-spark.html#pyspark-tests-with-maven-or-sbtCloses#27218 from MaxGekk/remove-respectSessionTimeZone.
Authored-by: Maxim Gekk <max.gekk@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
Change ```DecisionTreeClassifier``` to ```FMClassifier``` in ```OneVsRest``` setWeightCol test
### Why are the changes needed?
In ```OneVsRest```, if the classifier doesn't support instance weight, ```OneVsRest``` weightCol will be ignored, so unit test has tested one classifier(```LogisticRegression```) that support instance weight, and one classifier (```DecisionTreeClassifier```) that doesn't support instance weight. Since ```DecisionTreeClassifier``` now supports instance weight, we need to change it to the classifier that doesn't have weight support.
### Does this PR introduce any user-facing change?
No
### How was this patch tested?
Existing test
Closes#27204 from huaxingao/spark-ovr-minor.
Authored-by: Huaxin Gao <huaxing@us.ibm.com>
Signed-off-by: zhengruifeng <ruifengz@foxmail.com>
### What changes were proposed in this pull request?
add setInputCol/setOutputCol in OHEModel
### Why are the changes needed?
setInputCol/setOutputCol should be in OHEModel too.
### Does this PR introduce any user-facing change?
Yes.
```OHEModel.setInputCol```
```OHEModel.setOutputCol```
### How was this patch tested?
Manually tested.
Closes#27228 from huaxingao/spark-29565.
Authored-by: Huaxin Gao <huaxing@us.ibm.com>
Signed-off-by: zhengruifeng <ruifengz@foxmail.com>
### What changes were proposed in this pull request?
This is a followup of https://github.com/apache/spark/pull/27109. It should match the parameter lists in `createDataFrame`.
### Why are the changes needed?
To pass parameters supposed to pass.
### Does this PR introduce any user-facing change?
No (it's only in master)
### How was this patch tested?
Manually tested and existing tests should cover.
Closes#27225 from HyukjinKwon/SPARK-30434-followup.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
Removal of following `Param` fields:
- `factorSize`
- `fitLinear`
- `miniBatchFraction`
- `initStd`
- `solver`
from `FMClassifier` and `FMRegressor`
### Why are the changes needed?
This `Param` members are already provided by `_FactorizationMachinesParams`
0f3d744c3f/python/pyspark/ml/regression.py (L2303-L2318)
which is mixed into `FMRegressor`:
0f3d744c3f/python/pyspark/ml/regression.py (L2350)
and `FMClassifier`:
0f3d744c3f/python/pyspark/ml/classification.py (L2793)
### Does this PR introduce any user-facing change?
No
### How was this patch tested?
Manual testing.
Closes#27205 from zero323/SPARK-30378-FOLLOWUP.
Authored-by: zero323 <mszymkiewicz@gmail.com>
Signed-off-by: Sean Owen <srowen@gmail.com>
### What changes were proposed in this pull request?
This PR adjusts `_to_java` and `_from_java` of `OneVsRest` and `OneVsRestModel` to preserve `weightCol`.
### Why are the changes needed?
Currently both `Params` don't preserve `weightCol` `Params` when data is saved / loaded:
```python
from pyspark.ml.classification import LogisticRegression, OneVsRest, OneVsRestModel
from pyspark.ml.linalg import DenseVector
df = spark.createDataFrame([(0, 1, DenseVector([1.0, 0.0])), (0, 1, DenseVector([1.0, 0.0]))], ("label", "w", "features"))
ovr = OneVsRest(classifier=LogisticRegression()).setWeightCol("w")
ovrm = ovr.fit(df)
ovr.getWeightCol()
## 'w'
ovrm.getWeightCol()
## 'w'
ovr.write().overwrite().save("/tmp/ovr")
ovr_ = OneVsRest.load("/tmp/ovr")
ovr_.getWeightCol()
## KeyError
## ...
## KeyError: Param(parent='OneVsRest_5145d56b6bd1', name='weightCol', doc='weight column name. ...)
ovrm.write().overwrite().save("/tmp/ovrm")
ovrm_ = OneVsRestModel.load("/tmp/ovrm")
ovrm_ .getWeightCol()
## KeyError
## ...
## KeyError: Param(parent='OneVsRestModel_598c6d900fad', name='weightCol', doc='weight column name ...
```
### Does this PR introduce any user-facing change?
After this PR is merged, loaded objects will have `weightCol` `Param` set.
### How was this patch tested?
- Manual testing.
- Extension of existing persistence tests.
Closes#27190 from zero323/SPARK-30504.
Authored-by: zero323 <mszymkiewicz@gmail.com>
Signed-off-by: Sean Owen <srowen@gmail.com>