Disallow the use of unused imports:
- Unnecessary increases the memory footprint of the application
- Removes the imports that are required for the examples in the docstring from the file-scope to the example itself. This keeps the files itself clean, and gives a more complete example as it also includes the imports :)
```
fokkodriesprongFan spark % flake8 python | grep -i "imported but unused"
python/pyspark/cloudpickle.py:46:1: F401 'functools.partial' imported but unused
python/pyspark/cloudpickle.py:55:1: F401 'traceback' imported but unused
python/pyspark/heapq3.py:868:5: F401 '_heapq.*' imported but unused
python/pyspark/__init__.py:61:1: F401 'pyspark.version.__version__' imported but unused
python/pyspark/__init__.py:62:1: F401 'pyspark._globals._NoValue' imported but unused
python/pyspark/__init__.py:115:1: F401 'pyspark.sql.SQLContext' imported but unused
python/pyspark/__init__.py:115:1: F401 'pyspark.sql.HiveContext' imported but unused
python/pyspark/__init__.py:115:1: F401 'pyspark.sql.Row' imported but unused
python/pyspark/rdd.py:21:1: F401 're' imported but unused
python/pyspark/rdd.py:29:1: F401 'tempfile.NamedTemporaryFile' imported but unused
python/pyspark/mllib/regression.py:26:1: F401 'pyspark.mllib.linalg.SparseVector' imported but unused
python/pyspark/mllib/clustering.py:28:1: F401 'pyspark.mllib.linalg.SparseVector' imported but unused
python/pyspark/mllib/clustering.py:28:1: F401 'pyspark.mllib.linalg.DenseVector' imported but unused
python/pyspark/mllib/classification.py:26:1: F401 'pyspark.mllib.linalg.SparseVector' imported but unused
python/pyspark/mllib/feature.py:28:1: F401 'pyspark.mllib.linalg.DenseVector' imported but unused
python/pyspark/mllib/feature.py:28:1: F401 'pyspark.mllib.linalg.SparseVector' imported but unused
python/pyspark/mllib/feature.py:30:1: F401 'pyspark.mllib.regression.LabeledPoint' imported but unused
python/pyspark/mllib/tests/test_linalg.py:18:1: F401 'sys' imported but unused
python/pyspark/mllib/tests/test_linalg.py:642:5: F401 'pyspark.mllib.tests.test_linalg.*' imported but unused
python/pyspark/mllib/tests/test_feature.py:21:1: F401 'numpy.random' imported but unused
python/pyspark/mllib/tests/test_feature.py:21:1: F401 'numpy.exp' imported but unused
python/pyspark/mllib/tests/test_feature.py:23:1: F401 'pyspark.mllib.linalg.Vector' imported but unused
python/pyspark/mllib/tests/test_feature.py:23:1: F401 'pyspark.mllib.linalg.VectorUDT' imported but unused
python/pyspark/mllib/tests/test_feature.py:185:5: F401 'pyspark.mllib.tests.test_feature.*' imported but unused
python/pyspark/mllib/tests/test_util.py:97:5: F401 'pyspark.mllib.tests.test_util.*' imported but unused
python/pyspark/mllib/tests/test_stat.py:23:1: F401 'pyspark.mllib.linalg.Vector' imported but unused
python/pyspark/mllib/tests/test_stat.py:23:1: F401 'pyspark.mllib.linalg.SparseVector' imported but unused
python/pyspark/mllib/tests/test_stat.py:23:1: F401 'pyspark.mllib.linalg.DenseVector' imported but unused
python/pyspark/mllib/tests/test_stat.py:23:1: F401 'pyspark.mllib.linalg.VectorUDT' imported but unused
python/pyspark/mllib/tests/test_stat.py:23:1: F401 'pyspark.mllib.linalg._convert_to_vector' imported but unused
python/pyspark/mllib/tests/test_stat.py:23:1: F401 'pyspark.mllib.linalg.DenseMatrix' imported but unused
python/pyspark/mllib/tests/test_stat.py:23:1: F401 'pyspark.mllib.linalg.SparseMatrix' imported but unused
python/pyspark/mllib/tests/test_stat.py:23:1: F401 'pyspark.mllib.linalg.MatrixUDT' imported but unused
python/pyspark/mllib/tests/test_stat.py:181:5: F401 'pyspark.mllib.tests.test_stat.*' imported but unused
python/pyspark/mllib/tests/test_streaming_algorithms.py:18:1: F401 'time.time' imported but unused
python/pyspark/mllib/tests/test_streaming_algorithms.py:18:1: F401 'time.sleep' imported but unused
python/pyspark/mllib/tests/test_streaming_algorithms.py:470:5: F401 'pyspark.mllib.tests.test_streaming_algorithms.*' imported but unused
python/pyspark/mllib/tests/test_algorithms.py:295:5: F401 'pyspark.mllib.tests.test_algorithms.*' imported but unused
python/pyspark/tests/test_serializers.py:90:13: F401 'xmlrunner' imported but unused
python/pyspark/tests/test_rdd.py:21:1: F401 'sys' imported but unused
python/pyspark/tests/test_rdd.py:29:1: F401 'pyspark.resource.ResourceProfile' imported but unused
python/pyspark/tests/test_rdd.py:885:5: F401 'pyspark.tests.test_rdd.*' imported but unused
python/pyspark/tests/test_readwrite.py:19:1: F401 'sys' imported but unused
python/pyspark/tests/test_readwrite.py:22:1: F401 'array.array' imported but unused
python/pyspark/tests/test_readwrite.py:309:5: F401 'pyspark.tests.test_readwrite.*' imported but unused
python/pyspark/tests/test_join.py:62:5: F401 'pyspark.tests.test_join.*' imported but unused
python/pyspark/tests/test_taskcontext.py:19:1: F401 'shutil' imported but unused
python/pyspark/tests/test_taskcontext.py:325:5: F401 'pyspark.tests.test_taskcontext.*' imported but unused
python/pyspark/tests/test_conf.py:36:5: F401 'pyspark.tests.test_conf.*' imported but unused
python/pyspark/tests/test_broadcast.py:148:5: F401 'pyspark.tests.test_broadcast.*' imported but unused
python/pyspark/tests/test_daemon.py:76:5: F401 'pyspark.tests.test_daemon.*' imported but unused
python/pyspark/tests/test_util.py:77:5: F401 'pyspark.tests.test_util.*' imported but unused
python/pyspark/tests/test_pin_thread.py:19:1: F401 'random' imported but unused
python/pyspark/tests/test_pin_thread.py:149:5: F401 'pyspark.tests.test_pin_thread.*' imported but unused
python/pyspark/tests/test_worker.py:19:1: F401 'sys' imported but unused
python/pyspark/tests/test_worker.py:26:5: F401 'resource' imported but unused
python/pyspark/tests/test_worker.py:203:5: F401 'pyspark.tests.test_worker.*' imported but unused
python/pyspark/tests/test_profiler.py:101:5: F401 'pyspark.tests.test_profiler.*' imported but unused
python/pyspark/tests/test_shuffle.py:18:1: F401 'sys' imported but unused
python/pyspark/tests/test_shuffle.py:171:5: F401 'pyspark.tests.test_shuffle.*' imported but unused
python/pyspark/tests/test_rddbarrier.py:43:5: F401 'pyspark.tests.test_rddbarrier.*' imported but unused
python/pyspark/tests/test_context.py:129:13: F401 'userlibrary.UserClass' imported but unused
python/pyspark/tests/test_context.py:140:13: F401 'userlib.UserClass' imported but unused
python/pyspark/tests/test_context.py:310:5: F401 'pyspark.tests.test_context.*' imported but unused
python/pyspark/tests/test_appsubmit.py:241:5: F401 'pyspark.tests.test_appsubmit.*' imported but unused
python/pyspark/streaming/dstream.py:18:1: F401 'sys' imported but unused
python/pyspark/streaming/tests/test_dstream.py:27:1: F401 'pyspark.RDD' imported but unused
python/pyspark/streaming/tests/test_dstream.py:647:5: F401 'pyspark.streaming.tests.test_dstream.*' imported but unused
python/pyspark/streaming/tests/test_kinesis.py:83:5: F401 'pyspark.streaming.tests.test_kinesis.*' imported but unused
python/pyspark/streaming/tests/test_listener.py:152:5: F401 'pyspark.streaming.tests.test_listener.*' imported but unused
python/pyspark/streaming/tests/test_context.py:178:5: F401 'pyspark.streaming.tests.test_context.*' imported but unused
python/pyspark/testing/utils.py:30:5: F401 'scipy.sparse' imported but unused
python/pyspark/testing/utils.py:36:5: F401 'numpy as np' imported but unused
python/pyspark/ml/regression.py:25:1: F401 'pyspark.ml.tree._TreeEnsembleParams' imported but unused
python/pyspark/ml/regression.py:25:1: F401 'pyspark.ml.tree._HasVarianceImpurity' imported but unused
python/pyspark/ml/regression.py:29:1: F401 'pyspark.ml.wrapper.JavaParams' imported but unused
python/pyspark/ml/util.py:19:1: F401 'sys' imported but unused
python/pyspark/ml/__init__.py:25:1: F401 'pyspark.ml.pipeline' imported but unused
python/pyspark/ml/pipeline.py:18:1: F401 'sys' imported but unused
python/pyspark/ml/stat.py:22:1: F401 'pyspark.ml.linalg.DenseMatrix' imported but unused
python/pyspark/ml/stat.py:22:1: F401 'pyspark.ml.linalg.Vectors' imported but unused
python/pyspark/ml/tests/test_training_summary.py:18:1: F401 'sys' imported but unused
python/pyspark/ml/tests/test_training_summary.py:364:5: F401 'pyspark.ml.tests.test_training_summary.*' imported but unused
python/pyspark/ml/tests/test_linalg.py:381:5: F401 'pyspark.ml.tests.test_linalg.*' imported but unused
python/pyspark/ml/tests/test_tuning.py:427:9: F401 'pyspark.sql.functions as F' imported but unused
python/pyspark/ml/tests/test_tuning.py:757:5: F401 'pyspark.ml.tests.test_tuning.*' imported but unused
python/pyspark/ml/tests/test_wrapper.py:120:5: F401 'pyspark.ml.tests.test_wrapper.*' imported but unused
python/pyspark/ml/tests/test_feature.py:19:1: F401 'sys' imported but unused
python/pyspark/ml/tests/test_feature.py:304:5: F401 'pyspark.ml.tests.test_feature.*' imported but unused
python/pyspark/ml/tests/test_image.py:19:1: F401 'py4j' imported but unused
python/pyspark/ml/tests/test_image.py:22:1: F401 'pyspark.testing.mlutils.PySparkTestCase' imported but unused
python/pyspark/ml/tests/test_image.py:71:5: F401 'pyspark.ml.tests.test_image.*' imported but unused
python/pyspark/ml/tests/test_persistence.py:456:5: F401 'pyspark.ml.tests.test_persistence.*' imported but unused
python/pyspark/ml/tests/test_evaluation.py:56:5: F401 'pyspark.ml.tests.test_evaluation.*' imported but unused
python/pyspark/ml/tests/test_stat.py:43:5: F401 'pyspark.ml.tests.test_stat.*' imported but unused
python/pyspark/ml/tests/test_base.py:70:5: F401 'pyspark.ml.tests.test_base.*' imported but unused
python/pyspark/ml/tests/test_param.py:20:1: F401 'sys' imported but unused
python/pyspark/ml/tests/test_param.py:375:5: F401 'pyspark.ml.tests.test_param.*' imported but unused
python/pyspark/ml/tests/test_pipeline.py:62:5: F401 'pyspark.ml.tests.test_pipeline.*' imported but unused
python/pyspark/ml/tests/test_algorithms.py:333:5: F401 'pyspark.ml.tests.test_algorithms.*' imported but unused
python/pyspark/ml/param/__init__.py:18:1: F401 'sys' imported but unused
python/pyspark/resource/tests/test_resources.py:17:1: F401 'random' imported but unused
python/pyspark/resource/tests/test_resources.py:20:1: F401 'pyspark.resource.ResourceProfile' imported but unused
python/pyspark/resource/tests/test_resources.py:75:5: F401 'pyspark.resource.tests.test_resources.*' imported but unused
python/pyspark/sql/functions.py:32:1: F401 'pyspark.sql.udf.UserDefinedFunction' imported but unused
python/pyspark/sql/functions.py:34:1: F401 'pyspark.sql.pandas.functions.pandas_udf' imported but unused
python/pyspark/sql/session.py:30:1: F401 'pyspark.sql.types.Row' imported but unused
python/pyspark/sql/session.py:30:1: F401 'pyspark.sql.types.StringType' imported but unused
python/pyspark/sql/readwriter.py:1084:5: F401 'pyspark.sql.Row' imported but unused
python/pyspark/sql/context.py:26:1: F401 'pyspark.sql.types.IntegerType' imported but unused
python/pyspark/sql/context.py:26:1: F401 'pyspark.sql.types.Row' imported but unused
python/pyspark/sql/context.py:26:1: F401 'pyspark.sql.types.StringType' imported but unused
python/pyspark/sql/context.py:27:1: F401 'pyspark.sql.udf.UDFRegistration' imported but unused
python/pyspark/sql/streaming.py:1212:5: F401 'pyspark.sql.Row' imported but unused
python/pyspark/sql/tests/test_utils.py:55:5: F401 'pyspark.sql.tests.test_utils.*' imported but unused
python/pyspark/sql/tests/test_pandas_map.py:18:1: F401 'sys' imported but unused
python/pyspark/sql/tests/test_pandas_map.py:22:1: F401 'pyspark.sql.functions.pandas_udf' imported but unused
python/pyspark/sql/tests/test_pandas_map.py:22:1: F401 'pyspark.sql.functions.PandasUDFType' imported but unused
python/pyspark/sql/tests/test_pandas_map.py:119:5: F401 'pyspark.sql.tests.test_pandas_map.*' imported but unused
python/pyspark/sql/tests/test_catalog.py:193:5: F401 'pyspark.sql.tests.test_catalog.*' imported but unused
python/pyspark/sql/tests/test_group.py:39:5: F401 'pyspark.sql.tests.test_group.*' imported but unused
python/pyspark/sql/tests/test_session.py:361:5: F401 'pyspark.sql.tests.test_session.*' imported but unused
python/pyspark/sql/tests/test_conf.py:49:5: F401 'pyspark.sql.tests.test_conf.*' imported but unused
python/pyspark/sql/tests/test_pandas_cogrouped_map.py:19:1: F401 'sys' imported but unused
python/pyspark/sql/tests/test_pandas_cogrouped_map.py:21:1: F401 'pyspark.sql.functions.sum' imported but unused
python/pyspark/sql/tests/test_pandas_cogrouped_map.py:21:1: F401 'pyspark.sql.functions.PandasUDFType' imported but unused
python/pyspark/sql/tests/test_pandas_cogrouped_map.py:29:5: F401 'pandas.util.testing.assert_series_equal' imported but unused
python/pyspark/sql/tests/test_pandas_cogrouped_map.py:32:5: F401 'pyarrow as pa' imported but unused
python/pyspark/sql/tests/test_pandas_cogrouped_map.py:248:5: F401 'pyspark.sql.tests.test_pandas_cogrouped_map.*' imported but unused
python/pyspark/sql/tests/test_udf.py:24:1: F401 'py4j' imported but unused
python/pyspark/sql/tests/test_pandas_udf_typehints.py:246:5: F401 'pyspark.sql.tests.test_pandas_udf_typehints.*' imported but unused
python/pyspark/sql/tests/test_functions.py:19:1: F401 'sys' imported but unused
python/pyspark/sql/tests/test_functions.py:362:9: F401 'pyspark.sql.functions.exists' imported but unused
python/pyspark/sql/tests/test_functions.py:387:5: F401 'pyspark.sql.tests.test_functions.*' imported but unused
python/pyspark/sql/tests/test_pandas_udf_scalar.py:21:1: F401 'sys' imported but unused
python/pyspark/sql/tests/test_pandas_udf_scalar.py:45:5: F401 'pyarrow as pa' imported but unused
python/pyspark/sql/tests/test_pandas_udf_window.py:355:5: F401 'pyspark.sql.tests.test_pandas_udf_window.*' imported but unused
python/pyspark/sql/tests/test_arrow.py:38:5: F401 'pyarrow as pa' imported but unused
python/pyspark/sql/tests/test_pandas_grouped_map.py:20:1: F401 'sys' imported but unused
python/pyspark/sql/tests/test_pandas_grouped_map.py:38:5: F401 'pyarrow as pa' imported but unused
python/pyspark/sql/tests/test_dataframe.py:382:9: F401 'pyspark.sql.DataFrame' imported but unused
python/pyspark/sql/avro/functions.py:125:5: F401 'pyspark.sql.Row' imported but unused
python/pyspark/sql/pandas/functions.py:19:1: F401 'sys' imported but unused
```
After:
```
fokkodriesprongFan spark % flake8 python | grep -i "imported but unused"
fokkodriesprongFan spark %
```
### What changes were proposed in this pull request?
Removing unused imports from the Python files to keep everything nice and tidy.
### Why are the changes needed?
Cleaning up of the imports that aren't used, and suppressing the imports that are used as references to other modules, preserving backward compatibility.
### Does this PR introduce _any_ user-facing change?
No.
### How was this patch tested?
Adding the rule to the existing Flake8 checks.
Closes#29121 from Fokko/SPARK-32319.
Authored-by: Fokko Driesprong <fokko@apache.org>
Signed-off-by: Dongjoon Hyun <dongjoon@apache.org>
### What changes were proposed in this pull request?
This PR proposes to port the test case from https://github.com/apache/spark/pull/29098 to branch-3.0 and master. In the master and branch-3.0, this was fixed together at ecaa495b1f but no partition case is not being tested.
### Why are the changes needed?
To improve test coverage.
### Does this PR introduce _any_ user-facing change?
No, test-only.
### How was this patch tested?
Unit test was forward-ported.
Closes#29099 from HyukjinKwon/SPARK-32300-1.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR aims to drop Python 2.7, 3.4 and 3.5.
Roughly speaking, it removes all the widely known Python 2 compatibility workarounds such as `sys.version` comparison, `__future__`. Also, it removes the Python 2 dedicated codes such as `ArrayConstructor` in Spark.
### Why are the changes needed?
1. Unsupport EOL Python versions
2. Reduce maintenance overhead and remove a bit of legacy codes and hacks for Python 2.
3. PyPy2 has a critical bug that causes a flaky test, SPARK-28358 given my testing and investigation.
4. Users can use Python type hints with Pandas UDFs without thinking about Python version
5. Users can leverage one latest cloudpickle, https://github.com/apache/spark/pull/28950. With Python 3.8+ it can also leverage C pickle.
### Does this PR introduce _any_ user-facing change?
Yes, users cannot use Python 2.7, 3.4 and 3.5 in the upcoming Spark version.
### How was this patch tested?
Manually tested and also tested in Jenkins.
Closes#28957 from HyukjinKwon/SPARK-32138.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR aims to run the Spark tests in Github Actions.
To briefly explain the main idea:
- Reuse `dev/run-tests.py` with SBT build
- Reuse the modules in `dev/sparktestsupport/modules.py` to test each module
- Pass the modules to test into `dev/run-tests.py` directly via `TEST_ONLY_MODULES` environment variable. For example, `pyspark-sql,core,sql,hive`.
- `dev/run-tests.py` _does not_ take the dependent modules into account but solely the specified modules to test.
Another thing to note might be `SlowHiveTest` annotation. Running the tests in Hive modules takes too much so the slow tests are extracted and it runs as a separate job. It was extracted from the actual elapsed time in Jenkins:
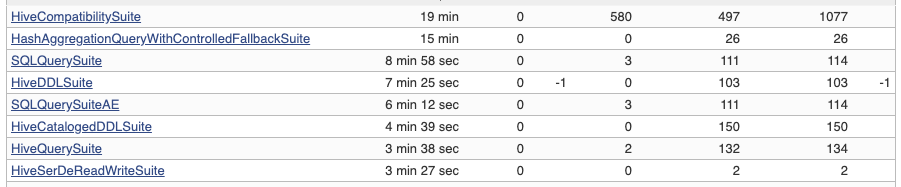
So, Hive tests are separated into to jobs. One is slow test cases, and the other one is the other test cases.
_Note that_ the current GitHub Actions build virtually copies what the default PR builder on Jenkins does (without other profiles such as JDK 11, Hadoop 2, etc.). The only exception is Kinesis https://github.com/apache/spark/pull/29057/files#diff-04eb107ee163a50b61281ca08f4e4c7bR23
### Why are the changes needed?
Last week and onwards, the Jenkins machines became very unstable for many reasons:
- Apparently, the machines became extremely slow. Almost all tests can't pass.
- One machine (worker 4) started to have the corrupt `.m2` which fails the build.
- Documentation build fails time to time for an unknown reason in Jenkins machine specifically. This is disabled for now at https://github.com/apache/spark/pull/29017.
- Almost all PRs are basically blocked by this instability currently.
The advantages of using Github Actions:
- To avoid depending on few persons who can access to the cluster.
- To reduce the elapsed time in the build - we could split the tests (e.g., SQL, ML, CORE), and run them in parallel so the total build time will significantly reduce.
- To control the environment more flexibly.
- Other contributors can test and propose to fix Github Actions configurations so we can distribute this build management cost.
Note that:
- The current build in Jenkins takes _more than 7 hours_. With Github actions it takes _less than 2 hours_
- We can now control the environments especially for Python easily.
- The test and build look more stable than the Jenkins'.
### Does this PR introduce _any_ user-facing change?
No, dev-only change.
### How was this patch tested?
Tested at https://github.com/HyukjinKwon/spark/pull/4Closes#29057 from HyukjinKwon/migrate-to-github-actions.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: Dongjoon Hyun <dongjoon@apache.org>
### What changes were proposed in this pull request?
When you use floats are index of pandas, it creates a Spark DataFrame with a wrong results as below when Arrow is enabled:
```bash
./bin/pyspark --conf spark.sql.execution.arrow.pyspark.enabled=true
```
```python
>>> import pandas as pd
>>> spark.createDataFrame(pd.DataFrame({'a': [1,2,3]}, index=[2., 3., 4.])).show()
+---+
| a|
+---+
| 1|
| 1|
| 2|
+---+
```
This is because direct slicing uses the value as index when the index contains floats:
```python
>>> pd.DataFrame({'a': [1,2,3]}, index=[2., 3., 4.])[2:]
a
2.0 1
3.0 2
4.0 3
>>> pd.DataFrame({'a': [1,2,3]}, index=[2., 3., 4.]).iloc[2:]
a
4.0 3
>>> pd.DataFrame({'a': [1,2,3]}, index=[2, 3, 4])[2:]
a
4 3
```
This PR proposes to explicitly use `iloc` to positionally slide when we create a DataFrame from a pandas DataFrame with Arrow enabled.
FWIW, I was trying to investigate why direct slicing refers the index value or the positional index sometimes but I stopped investigating further after reading this https://pandas.pydata.org/pandas-docs/stable/getting_started/10min.html#selection
> While standard Python / Numpy expressions for selecting and setting are intuitive and come in handy for interactive work, for production code, we recommend the optimized pandas data access methods, `.at`, `.iat`, `.loc` and `.iloc`.
### Why are the changes needed?
To create the correct Spark DataFrame from a pandas DataFrame without a data loss.
### Does this PR introduce _any_ user-facing change?
Yes, it is a bug fix.
```bash
./bin/pyspark --conf spark.sql.execution.arrow.pyspark.enabled=true
```
```python
import pandas as pd
spark.createDataFrame(pd.DataFrame({'a': [1,2,3]}, index=[2., 3., 4.])).show()
```
Before:
```
+---+
| a|
+---+
| 1|
| 1|
| 2|
+---+
```
After:
```
+---+
| a|
+---+
| 1|
| 2|
| 3|
+---+
```
### How was this patch tested?
Manually tested and unittest were added.
Closes#28928 from HyukjinKwon/SPARK-32098.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: Bryan Cutler <cutlerb@gmail.com>
### What changes were proposed in this pull request?
Followup to make assertions from recent test consistent with the rest of the module
### Why are the changes needed?
Better to use assertions from `unittest` and be consistent
### Does this PR introduce _any_ user-facing change?
No
### How was this patch tested?
Existing tests
Closes#28659 from BryanCutler/arrow-category-test-fix-SPARK-25351.
Authored-by: Bryan Cutler <cutlerb@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
Handle Pandas category type while converting from python with Arrow enabled. The category column will be converted to whatever type the category elements are as is the case with Arrow disabled.
### Does this PR introduce any user-facing change?
No
### How was this patch tested?
New unit tests were added for `createDataFrame` and scalar `pandas_udf`
Closes#26585 from jalpan-randeri/feature-pyarrow-dictionary-type.
Authored-by: Jalpan Randeri <randerij@amazon.com>
Signed-off-by: Bryan Cutler <cutlerb@gmail.com>
### What changes were proposed in this pull request?
Fix PySpark test failures for using Pandas >= 1.0.0.
### Why are the changes needed?
Pandas 1.0.0 has recently been released and has API changes that result in PySpark test failures, this PR fixes the broken tests.
### Does this PR introduce any user-facing change?
No
### How was this patch tested?
Manually tested with Pandas 1.0.1 and PyArrow 0.16.0
Closes#27529 from BryanCutler/pandas-fix-tests-1.0-SPARK-30777.
Authored-by: Bryan Cutler <cutlerb@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
In the PR, I propose to remove the SQL config `spark.sql.execution.pandas.respectSessionTimeZone` which has been deprecated since Spark 2.3.
### Why are the changes needed?
To improve code maintainability.
### Does this PR introduce any user-facing change?
Yes.
### How was this patch tested?
by running python tests, https://spark.apache.org/docs/latest/building-spark.html#pyspark-tests-with-maven-or-sbtCloses#27218 from MaxGekk/remove-respectSessionTimeZone.
Authored-by: Maxim Gekk <max.gekk@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR proposes to move pandas related functionalities into pandas package. Namely:
```bash
pyspark/sql/pandas
├── __init__.py
├── conversion.py # Conversion between pandas <> PySpark DataFrames
├── functions.py # pandas_udf
├── group_ops.py # Grouped UDF / Cogrouped UDF + groupby.apply, groupby.cogroup.apply
├── map_ops.py # Map Iter UDF + mapInPandas
├── serializers.py # pandas <> PyArrow serializers
├── types.py # Type utils between pandas <> PyArrow
└── utils.py # Version requirement checks
```
In order to separately locate `groupby.apply`, `groupby.cogroup.apply`, `mapInPandas`, `toPandas`, and `createDataFrame(pdf)` under `pandas` sub-package, I had to use a mix-in approach which Scala side uses often by `trait`, and also pandas itself uses this approach (see `IndexOpsMixin` as an example) to group related functionalities. Currently, you can think it's like Scala's self typed trait. See the structure below:
```python
class PandasMapOpsMixin(object):
def mapInPandas(self, ...):
...
return ...
# other Pandas <> PySpark APIs
```
```python
class DataFrame(PandasMapOpsMixin):
# other DataFrame APIs equivalent to Scala side.
```
Yes, This is a big PR but they are mostly just moving around except one case `createDataFrame` which I had to split the methods.
### Why are the changes needed?
There are pandas functionalities here and there and I myself gets lost where it was. Also, when you have to make a change commonly for all of pandas related features, it's almost impossible now.
Also, after this change, `DataFrame` and `SparkSession` become more consistent with Scala side since pandas is specific to Python, and this change separates pandas-specific APIs away from `DataFrame` or `SparkSession`.
### Does this PR introduce any user-facing change?
No.
### How was this patch tested?
Existing tests should cover. Also, I manually built the PySpark API documentation and checked.
Closes#27109 from HyukjinKwon/pandas-refactoring.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR proposes to match the test with branch-2.4. See https://github.com/apache/spark/pull/25593#discussion_r318109047
Seems using `SparkSession.builder` with Spark conf possibly affects other tests.
### Why are the changes needed?
To match with branch-2.4 and to make easier to backport.
### Does this PR introduce any user-facing change?
No.
### How was this patch tested?
Test was fixed.
Closes#25603 from HyukjinKwon/SPARK-28881-followup.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
### What changes were proposed in this pull request?
This PR proposes to add a test case for:
```bash
./bin/pyspark --conf spark.driver.maxResultSize=1m
spark.conf.set("spark.sql.execution.arrow.enabled",True)
```
```python
spark.range(10000000).toPandas()
```
```
Empty DataFrame
Columns: [id]
Index: []
```
which can result in partial results (see https://github.com/apache/spark/pull/25593#issuecomment-525153808). This regression was found between Spark 2.3 and Spark 2.4, and accidentally fixed.
### Why are the changes needed?
To prevent the same regression in the future.
### Does this PR introduce any user-facing change?
No.
### How was this patch tested?
Test was added.
Closes#25594 from HyukjinKwon/SPARK-28881.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
In Python 2.7 with latest PyArrow and Pandas, the error message seems a bit different with Python 3. This PR simply fixes the test.
```
======================================================================
FAIL: test_createDataFrame_with_incorrect_schema (pyspark.sql.tests.test_arrow.ArrowTests)
----------------------------------------------------------------------
Traceback (most recent call last):
File "/.../spark/python/pyspark/sql/tests/test_arrow.py", line 275, in test_createDataFrame_with_incorrect_schema
self.spark.createDataFrame(pdf, schema=wrong_schema)
AssertionError: "integer.*required.*got.*str" does not match "('Exception thrown when converting pandas.Series (object) to Arrow Array (int32). It can be caused by overflows or other unsafe conversions warned by Arrow. Arrow safe type check can be disabled by using SQL config `spark.sql.execution.pandas.arrowSafeTypeConversion`.', ArrowTypeError('an integer is required',))"
======================================================================
FAIL: test_createDataFrame_with_incorrect_schema (pyspark.sql.tests.test_arrow.EncryptionArrowTests)
----------------------------------------------------------------------
Traceback (most recent call last):
File "/.../spark/python/pyspark/sql/tests/test_arrow.py", line 275, in test_createDataFrame_with_incorrect_schema
self.spark.createDataFrame(pdf, schema=wrong_schema)
AssertionError: "integer.*required.*got.*str" does not match "('Exception thrown when converting pandas.Series (object) to Arrow Array (int32). It can be caused by overflows or other unsafe conversions warned by Arrow. Arrow safe type check can be disabled by using SQL config `spark.sql.execution.pandas.arrowSafeTypeConversion`.', ArrowTypeError('an integer is required',))"
```
## How was this patch tested?
Manually tested.
```
cd python
./run-tests --python-executables=python --modules pyspark-sql
```
Closes#25042 from HyukjinKwon/SPARK-28240.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
Currently with `toLocalIterator()` and `toPandas()` with Arrow enabled, if the Spark job being run in the background serving thread errors, it will be caught and sent to Python through the PySpark serializer.
This is not the ideal solution because it is only catch a SparkException, it won't handle an error that occurs in the serializer, and each method has to have it's own special handling to propagate the error.
This PR instead returns the Python Server object along with the serving port and authentication info, so that it allows the Python caller to join with the serving thread. During the call to join, the serving thread Future is completed either successfully or with an exception. In the latter case, the exception will be propagated to Python through the Py4j call.
## How was this patch tested?
Existing tests
Closes#24834 from BryanCutler/pyspark-propagate-server-error-SPARK-27992.
Authored-by: Bryan Cutler <cutlerb@gmail.com>
Signed-off-by: Bryan Cutler <cutlerb@gmail.com>
## What changes were proposed in this pull request?
This patch removes `fillna(0)` when creating ArrowBatch from a pandas Series.
With `fillna(0)`, the original code would turn a timestamp type into object type, which pyarrow will complain later:
```
>>> s = pd.Series([pd.NaT, pd.Timestamp('2015-01-01')])
>>> s.dtypes
dtype('<M8[ns]')
>>> s.fillna(0)
0 0
1 2015-01-01 00:00:00
dtype: object
```
## How was this patch tested?
Added `test_timestamp_nat`
Closes#24844 from icexelloss/SPARK-28003-arrow-nat.
Authored-by: Li Jin <ice.xelloss@gmail.com>
Signed-off-by: Bryan Cutler <cutlerb@gmail.com>
## What changes were proposed in this pull request?
Currently, pretty skipped message added by f7435bec6a mechanism seems not working when xmlrunner is installed apparently.
This PR fixes two things:
1. When `xmlrunner` is installed, seems `xmlrunner` does not respect `vervosity` level in unittests (default is level 1).
So the output looks as below
```
Running tests...
----------------------------------------------------------------------
SSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSS
----------------------------------------------------------------------
```
So it is not caught by our message detection mechanism.
2. If we manually set the `vervocity` level to `xmlrunner`, it prints messages as below:
```
test_mixed_udf (pyspark.sql.tests.test_pandas_udf_scalar.ScalarPandasUDFTests) ... SKIP (0.000s)
test_mixed_udf_and_sql (pyspark.sql.tests.test_pandas_udf_scalar.ScalarPandasUDFTests) ... SKIP (0.000s)
...
```
This is different in our Jenkins machine:
```
test_createDataFrame_column_name_encoding (pyspark.sql.tests.test_arrow.ArrowTests) ... skipped 'Pandas >= 0.23.2 must be installed; however, it was not found.'
test_createDataFrame_does_not_modify_input (pyspark.sql.tests.test_arrow.ArrowTests) ... skipped 'Pandas >= 0.23.2 must be installed; however, it was not found.'
...
```
Note that last `SKIP` is different. This PR fixes the regular expression to catch `SKIP` case as well.
## How was this patch tested?
Manually tested.
**Before:**
```
Starting test(python2.7): pyspark....
Finished test(python2.7): pyspark.... (0s)
...
Tests passed in 562 seconds
========================================================================
...
```
**After:**
```
Starting test(python2.7): pyspark....
Finished test(python2.7): pyspark.... (48s) ... 93 tests were skipped
...
Tests passed in 560 seconds
Skipped tests pyspark.... with python2.7:
pyspark...(...) ... SKIP (0.000s)
...
========================================================================
...
```
Closes#24927 from HyukjinKwon/SPARK-28130.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
This increases the minimum supported version of Pandas to 0.23.2. Using a lower version will raise an error `Pandas >= 0.23.2 must be installed; however, your version was 0.XX`. Also, a workaround for using pyarrow with Pandas 0.19.2 was removed.
## How was this patch tested?
Existing Tests
Closes#24867 from BryanCutler/pyspark-increase-min-pandas-SPARK-28041.
Authored-by: Bryan Cutler <cutlerb@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
Similar to https://github.com/apache/spark/pull/24070, we now propagate SparkExceptions that are encountered during the collect in the java process to the python process.
Fixes https://jira.apache.org/jira/browse/SPARK-27805
## How was this patch tested?
Added a new unit test
Closes#24677 from dvogelbacher/dv/betterErrorMsgWhenUsingArrow.
Authored-by: David Vogelbacher <dvogelbacher@palantir.com>
Signed-off-by: Bryan Cutler <cutlerb@gmail.com>
## What changes were proposed in this pull request?
`spark.sql.execution.arrow.enabled` was added when we add PySpark arrow optimization.
Later, in the current master, SparkR arrow optimization was added and it's controlled by the same configuration `spark.sql.execution.arrow.enabled`.
There look two issues about this:
1. `spark.sql.execution.arrow.enabled` in PySpark was added from 2.3.0 whereas SparkR optimization was added 3.0.0. The stability is different so it's problematic when we change the default value for one of both optimization first.
2. Suppose users want to share some JVM by PySpark and SparkR. They are currently forced to use the optimization for all or none if the configuration is set globally.
This PR proposes two separate configuration groups for PySpark and SparkR about Arrow optimization:
- Deprecate `spark.sql.execution.arrow.enabled`
- Add `spark.sql.execution.arrow.pyspark.enabled` (fallback to `spark.sql.execution.arrow.enabled`)
- Add `spark.sql.execution.arrow.sparkr.enabled`
- Deprecate `spark.sql.execution.arrow.fallback.enabled`
- Add `spark.sql.execution.arrow.pyspark.fallback.enabled ` (fallback to `spark.sql.execution.arrow.fallback.enabled`)
Note that `spark.sql.execution.arrow.maxRecordsPerBatch` is used within JVM side for both.
Note that `spark.sql.execution.arrow.fallback.enabled` was added due to behaviour change. We don't need it in SparkR - SparkR side has the automatic fallback.
## How was this patch tested?
Manually tested and some unittests were added.
Closes#24700 from HyukjinKwon/separate-sparkr-arrow.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
https://github.com/apache/spark/pull/22275 introduced a performance improvement where we send partitions out of order to python and then, as a last step, send the partition order as well.
However, if there are no partitions we will never send the partition order and we will get an "EofError" on the python side.
This PR fixes this by also sending the partition order if there are no partitions present.
## How was this patch tested?
New unit test added.
Closes#24650 from dvogelbacher/dv/fixNoPartitionArrowConversion.
Authored-by: David Vogelbacher <dvogelbacher@palantir.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
This increases the minimum support version of pyarrow to 0.12.1 and removes workarounds in pyspark to remain compatible with prior versions. This means that users will need to have at least pyarrow 0.12.1 installed and available in the cluster or an `ImportError` will be raised to indicate an upgrade is needed.
## How was this patch tested?
Existing tests using:
Python 2.7.15, pyarrow 0.12.1, pandas 0.24.2
Python 3.6.7, pyarrow 0.12.1, pandas 0.24.0
Closes#24298 from BryanCutler/arrow-bump-min-pyarrow-SPARK-27276.
Authored-by: Bryan Cutler <cutlerb@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
Running PySpark tests with Pandas 0.24.x causes a failure in `test_pandas_udf_grouped_map` test_supported_types:
`ValueError: The truth value of an array with more than one element is ambiguous. Use a.any() or a.all()`
This is because a column is an ArrayType and the method `sqlutils ReusedSQLTestCase.assertPandasEqual ` does not properly check this.
This PR removes `assertPandasEqual` and replaces it with the built-in `pandas.util.testing.assert_frame_equal` which can properly handle columns of ArrayType and also prints out better diff between the DataFrames when an error occurs.
Additionally, imports of pandas and pyarrow were moved to the top of related test files to avoid duplicating the same import many times.
## How was this patch tested?
Existing tests
Closes#24306 from BryanCutler/python-pandas-assert_frame_equal-SPARK-27387.
Authored-by: Bryan Cutler <cutlerb@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
Currently `DataFrame.toPandas()` with arrow enabled or `ArrowStreamPandasSerializer` for pandas UDF with pyarrow<0.12 creates `datetime64[ns]` type series as intermediate data and then convert to `datetime.date` series, but the intermediate `datetime64[ns]` might cause an overflow even if the date is valid.
```
>>> import datetime
>>>
>>> t = [datetime.date(2262, 4, 12), datetime.date(2263, 4, 12)]
>>>
>>> df = spark.createDataFrame(t, 'date')
>>> df.show()
+----------+
| value|
+----------+
|2262-04-12|
|2263-04-12|
+----------+
>>>
>>> spark.conf.set("spark.sql.execution.arrow.enabled", "true")
>>>
>>> df.toPandas()
value
0 1677-09-21
1 1678-09-21
```
We should avoid creating such intermediate data and create `datetime.date` series directly instead.
## How was this patch tested?
Modified some tests to include the date which overflow caused by the intermediate conversion.
Run tests with pyarrow 0.8, 0.10, 0.11, 0.12 in my local environment.
Closes#23795 from ueshin/issues/SPARK-26887/date_as_object.
Authored-by: Takuya UESHIN <ueshin@databricks.com>
Signed-off-by: Hyukjin Kwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
Upgrade Apache Arrow to version 0.12.0. This includes the Java artifacts and fixes to enable usage with pyarrow 0.12.0
Version 0.12.0 includes the following selected fixes/improvements relevant to Spark users:
* Safe cast fails from numpy float64 array with nans to integer, ARROW-4258
* Java, Reduce heap usage for variable width vectors, ARROW-4147
* Binary identity cast not implemented, ARROW-4101
* pyarrow open_stream deprecated, use ipc.open_stream, ARROW-4098
* conversion to date object no longer needed, ARROW-3910
* Error reading IPC file with no record batches, ARROW-3894
* Signed to unsigned integer cast yields incorrect results when type sizes are the same, ARROW-3790
* from_pandas gives incorrect results when converting floating point to bool, ARROW-3428
* Import pyarrow fails if scikit-learn is installed from conda (boost-cpp / libboost issue), ARROW-3048
* Java update to official Flatbuffers version 1.9.0, ARROW-3175
complete list [here](https://issues.apache.org/jira/issues/?jql=project%20%3D%20ARROW%20AND%20status%20in%20(Resolved%2C%20Closed)%20AND%20fixVersion%20%3D%200.12.0)
PySpark requires the following fixes to work with PyArrow 0.12.0
* Encrypted pyspark worker fails due to ChunkedStream missing closed property
* pyarrow now converts dates as objects by default, which causes error because type is assumed datetime64
* ArrowTests fails due to difference in raised error message
* pyarrow.open_stream deprecated
* tests fail because groupby adds index column with duplicate name
## How was this patch tested?
Ran unit tests with pyarrow versions 0.8.0, 0.10.0, 0.11.1, 0.12.0
Closes#23657 from BryanCutler/arrow-upgrade-012.
Authored-by: Bryan Cutler <cutlerb@gmail.com>
Signed-off-by: Hyukjin Kwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
When executing `toPandas` with Arrow enabled, partitions that arrive in the JVM out-of-order must be buffered before they can be send to Python. This causes an excess of memory to be used in the driver JVM and increases the time it takes to complete because data must sit in the JVM waiting for preceding partitions to come in.
This change sends un-ordered partitions to Python as soon as they arrive in the JVM, followed by a list of partition indices so that Python can assemble the data in the correct order. This way, data is not buffered at the JVM and there is no waiting on particular partitions so performance will be increased.
Followup to #21546
## How was this patch tested?
Added new test with a large number of batches per partition, and test that forces a small delay in the first partition. These test that partitions are collected out-of-order and then are are put in the correct order in Python.
## Performance Tests - toPandas
Tests run on a 4 node standalone cluster with 32 cores total, 14.04.1-Ubuntu and OpenJDK 8
measured wall clock time to execute `toPandas()` and took the average best time of 5 runs/5 loops each.
Test code
```python
df = spark.range(1 << 25, numPartitions=32).toDF("id").withColumn("x1", rand()).withColumn("x2", rand()).withColumn("x3", rand()).withColumn("x4", rand())
for i in range(5):
start = time.time()
_ = df.toPandas()
elapsed = time.time() - start
```
Spark config
```
spark.driver.memory 5g
spark.executor.memory 5g
spark.driver.maxResultSize 2g
spark.sql.execution.arrow.enabled true
```
Current Master w/ Arrow stream | This PR
---------------------|------------
5.16207 | 4.342533
5.133671 | 4.399408
5.147513 | 4.468471
5.105243 | 4.36524
5.018685 | 4.373791
Avg Master | Avg This PR
------------------|--------------
5.1134364 | 4.3898886
Speedup of **1.164821449**
Closes#22275 from BryanCutler/arrow-toPandas-oo-batches-SPARK-25274.
Authored-by: Bryan Cutler <cutlerb@gmail.com>
Signed-off-by: Bryan Cutler <cutlerb@gmail.com>
## What changes were proposed in this pull request?
This PR continues to break down a big large file into smaller files. See https://github.com/apache/spark/pull/23021. It targets to follow https://github.com/numpy/numpy/tree/master/numpy.
Basically this PR proposes to break down `pyspark/tests.py` into ...:
```
pyspark
...
├── testing
...
│ └── utils.py
├── tests
│ ├── __init__.py
│ ├── test_appsubmit.py
│ ├── test_broadcast.py
│ ├── test_conf.py
│ ├── test_context.py
│ ├── test_daemon.py
│ ├── test_join.py
│ ├── test_profiler.py
│ ├── test_rdd.py
│ ├── test_readwrite.py
│ ├── test_serializers.py
│ ├── test_shuffle.py
│ ├── test_taskcontext.py
│ ├── test_util.py
│ └── test_worker.py
...
```
## How was this patch tested?
Existing tests should cover.
`cd python` and .`/run-tests-with-coverage`. Manually checked they are actually being ran.
Each test (not officially) can be ran via:
```bash
SPARK_TESTING=1 ./bin/pyspark pyspark.tests.test_context
```
Note that if you're using Mac and Python 3, you might have to `OBJC_DISABLE_INITIALIZE_FORK_SAFETY=YES`.
Closes#23033 from HyukjinKwon/SPARK-26036.
Authored-by: hyukjinkwon <gurwls223@apache.org>
Signed-off-by: hyukjinkwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
This is the official first attempt to break huge single `tests.py` file - I did it locally before few times and gave up for some reasons. Now, currently it really makes the unittests super hard to read and difficult to check. To me, it even bothers me to to scroll down the big file. It's one single 7000 lines file!
This is not only readability issue. Since one big test takes most of tests time, the tests don't run in parallel fully - although it will costs to start and stop the context.
We could pick up one example and follow. Given my investigation, the current style looks closer to NumPy structure and looks easier to follow. Please see https://github.com/numpy/numpy/tree/master/numpy.
Basically this PR proposes to break down `pyspark/sql/tests.py` into ...:
```bash
pyspark
...
├── sql
...
│ ├── tests # Includes all tests broken down from 'pyspark/sql/tests.py'
│ │ │ # Each matchs to module in 'pyspark/sql'. Additionally, some logical group can
│ │ │ # be added. For instance, 'test_arrow.py', 'test_datasources.py' ...
│ │ ├── __init__.py
│ │ ├── test_appsubmit.py
│ │ ├── test_arrow.py
│ │ ├── test_catalog.py
│ │ ├── test_column.py
│ │ ├── test_conf.py
│ │ ├── test_context.py
│ │ ├── test_dataframe.py
│ │ ├── test_datasources.py
│ │ ├── test_functions.py
│ │ ├── test_group.py
│ │ ├── test_pandas_udf.py
│ │ ├── test_pandas_udf_grouped_agg.py
│ │ ├── test_pandas_udf_grouped_map.py
│ │ ├── test_pandas_udf_scalar.py
│ │ ├── test_pandas_udf_window.py
│ │ ├── test_readwriter.py
│ │ ├── test_serde.py
│ │ ├── test_session.py
│ │ ├── test_streaming.py
│ │ ├── test_types.py
│ │ ├── test_udf.py
│ │ └── test_utils.py
...
├── testing # Includes testing utils that can be used in unittests.
│ ├── __init__.py
│ └── sqlutils.py
...
```
## How was this patch tested?
Existing tests should cover.
`cd python` and `./run-tests-with-coverage`. Manually checked they are actually being ran.
Each test (not officially) can be ran via:
```
SPARK_TESTING=1 ./bin/pyspark pyspark.sql.tests.test_pandas_udf_scalar
```
Note that if you're using Mac and Python 3, you might have to `OBJC_DISABLE_INITIALIZE_FORK_SAFETY=YES`.
Closes#23021 from HyukjinKwon/SPARK-25344.
Authored-by: hyukjinkwon <gurwls223@apache.org>
Signed-off-by: hyukjinkwon <gurwls223@apache.org>