## What changes were proposed in this pull request?
This upgraded to a newer version of Pyrolite. Most updates [1] in the newer version are for dotnot. For java, it includes a bug fix to Unpickler regarding cleaning up Unpickler memo, and support of protocol 5.
After upgrading, we can remove the fix at SPARK-27629 for the bug in Unpickler.
[1] https://github.com/irmen/Pyrolite/compare/pyrolite-4.23...master
## How was this patch tested?
Manually tested on Python 3.6 in local on existing tests.
Closes#25143 from viirya/upgrade-pyrolite.
Authored-by: Liang-Chi Hsieh <viirya@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
The `_prepare_for_python_RDD` method currently broadcasts a pickled command if its length is greater than the hardcoded value `1 << 20` (1M). This change sets this value as a Spark conf instead.
## How was this patch tested?
Unit tests, manual tests.
Closes#25123 from jessecai/SPARK-28355.
Authored-by: Jesse Cai <jesse.cai@databricks.com>
Signed-off-by: gatorsmile <gatorsmile@gmail.com>
## What changes were proposed in this pull request?
Add python api support and JavaSparkContext support for resources(). I needed the JavaSparkContext support for it to properly translate into python with the py4j stuff.
## How was this patch tested?
Unit tests added and manually tested in local cluster mode and on yarn.
Closes#25087 from tgravescs/SPARK-28234-python.
Authored-by: Thomas Graves <tgraves@nvidia.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
There is a bug in `ExtractPythonUDFs` that produces wrong result attributes. It causes a failure when using `PythonUDF`s among multiple child plans, e.g., join. An example is using `PythonUDF`s in join condition.
```python
>>> left = spark.createDataFrame([Row(a=1, a1=1, a2=1), Row(a=2, a1=2, a2=2)])
>>> right = spark.createDataFrame([Row(b=1, b1=1, b2=1), Row(b=1, b1=3, b2=1)])
>>> f = udf(lambda a: a, IntegerType())
>>> df = left.join(right, [f("a") == f("b"), left.a1 == right.b1])
>>> df.collect()
19/07/10 12:20:49 ERROR Executor: Exception in task 5.0 in stage 0.0 (TID 5)
java.lang.ArrayIndexOutOfBoundsException: 1
at org.apache.spark.sql.catalyst.expressions.GenericInternalRow.genericGet(rows.scala:201)
at org.apache.spark.sql.catalyst.expressions.BaseGenericInternalRow.getAs(rows.scala:35)
at org.apache.spark.sql.catalyst.expressions.BaseGenericInternalRow.isNullAt(rows.scala:36)
at org.apache.spark.sql.catalyst.expressions.BaseGenericInternalRow.isNullAt$(rows.scala:36)
at org.apache.spark.sql.catalyst.expressions.GenericInternalRow.isNullAt(rows.scala:195)
at org.apache.spark.sql.catalyst.expressions.JoinedRow.isNullAt(JoinedRow.scala:70)
...
```
## How was this patch tested?
Added test.
Closes#25091 from viirya/SPARK-28323.
Authored-by: Liang-Chi Hsieh <viirya@gmail.com>
Signed-off-by: Bryan Cutler <cutlerb@gmail.com>
## What changes were proposed in this pull request?
In both cases, the input `DataFrame` schema must contain only the information that's required for the matrix object, so a vector column in the case of `RowMatrix` and long and vector columns for `IndexedRowMatrix`.
## How was this patch tested?
Unit tests that verify:
- `RowMatrix` and `IndexedRowMatrix` can be created from `DataFrame`s
- If the schema does not match expectations, we throw an `IllegalArgumentException`
Please review https://spark.apache.org/contributing.html before opening a pull request.
Closes#24953 from henrydavidge/row-matrix-df.
Authored-by: Henry D <henrydavidge@gmail.com>
Signed-off-by: Sean Owen <sean.owen@databricks.com>
## What changes were proposed in this pull request?
This PR adds some tests converted from `pgSQL/case.sql'` to test UDFs. Please see contribution guide of this umbrella ticket - [SPARK-27921](https://issues.apache.org/jira/browse/SPARK-27921).
This PR also contains two minor fixes:
1. Change name of Scala UDF from `UDF:name(...)` to `name(...)` to be consistent with Python'
2. Fix Scala UDF at `IntegratedUDFTestUtils.scala ` to handle `null` in strings.
<details><summary>Diff comparing to 'pgSQL/case.sql'</summary>
<p>
```diff
diff --git a/sql/core/src/test/resources/sql-tests/results/pgSQL/case.sql.out b/sql/core/src/test/resources/sql-tests/results/udf/pgSQL/udf-case.sql.out
index fa078d16d6d..55bef64338f 100644
--- a/sql/core/src/test/resources/sql-tests/results/pgSQL/case.sql.out
+++ b/sql/core/src/test/resources/sql-tests/results/udf/pgSQL/udf-case.sql.out
-115,7 +115,7 struct<>
-- !query 13
SELECT '3' AS `One`,
CASE
- WHEN 1 < 2 THEN 3
+ WHEN CAST(udf(1 < 2) AS boolean) THEN 3
END AS `Simple WHEN`
-- !query 13 schema
struct<One:string,Simple WHEN:int>
-126,10 +126,10 struct<One:string,Simple WHEN:int>
-- !query 14
SELECT '<NULL>' AS `One`,
CASE
- WHEN 1 > 2 THEN 3
+ WHEN 1 > 2 THEN udf(3)
END AS `Simple default`
-- !query 14 schema
-struct<One:string,Simple default:int>
+struct<One:string,Simple default:string>
-- !query 14 output
<NULL> NULL
-137,17 +137,17 struct<One:string,Simple default:int>
-- !query 15
SELECT '3' AS `One`,
CASE
- WHEN 1 < 2 THEN 3
- ELSE 4
+ WHEN udf(1) < 2 THEN udf(3)
+ ELSE udf(4)
END AS `Simple ELSE`
-- !query 15 schema
-struct<One:string,Simple ELSE:int>
+struct<One:string,Simple ELSE:string>
-- !query 15 output
3 3
-- !query 16
-SELECT '4' AS `One`,
+SELECT udf('4') AS `One`,
CASE
WHEN 1 > 2 THEN 3
ELSE 4
-159,10 +159,10 struct<One:string,ELSE default:int>
-- !query 17
-SELECT '6' AS `One`,
+SELECT udf('6') AS `One`,
CASE
- WHEN 1 > 2 THEN 3
- WHEN 4 < 5 THEN 6
+ WHEN CAST(udf(1 > 2) AS boolean) THEN 3
+ WHEN udf(4) < 5 THEN 6
ELSE 7
END AS `Two WHEN with default`
-- !query 17 schema
-173,7 +173,7 struct<One:string,Two WHEN with default:int>
-- !query 18
SELECT '7' AS `None`,
- CASE WHEN rand() < 0 THEN 1
+ CASE WHEN rand() < udf(0) THEN 1
END AS `NULL on no matches`
-- !query 18 schema
struct<None:string,NULL on no matches:int>
-182,36 +182,36 struct<None:string,NULL on no matches:int>
-- !query 19
-SELECT CASE WHEN 1=0 THEN 1/0 WHEN 1=1 THEN 1 ELSE 2/0 END
+SELECT CASE WHEN CAST(udf(1=0) AS boolean) THEN 1/0 WHEN 1=1 THEN 1 ELSE 2/0 END
-- !query 19 schema
-struct<CASE WHEN (1 = 0) THEN (CAST(1 AS DOUBLE) / CAST(0 AS DOUBLE)) WHEN (1 = 1) THEN CAST(1 AS DOUBLE) ELSE (CAST(2 AS DOUBLE) / CAST(0 AS DOUBLE)) END:double>
+struct<CASE WHEN CAST(udf((1 = 0)) AS BOOLEAN) THEN (CAST(1 AS DOUBLE) / CAST(0 AS DOUBLE)) WHEN (1 = 1) THEN CAST(1 AS DOUBLE) ELSE (CAST(2 AS DOUBLE) / CAST(0 AS DOUBLE)) END:double>
-- !query 19 output
1.0
-- !query 20
-SELECT CASE 1 WHEN 0 THEN 1/0 WHEN 1 THEN 1 ELSE 2/0 END
+SELECT CASE 1 WHEN 0 THEN 1/udf(0) WHEN 1 THEN 1 ELSE 2/0 END
-- !query 20 schema
-struct<CASE WHEN (1 = 0) THEN (CAST(1 AS DOUBLE) / CAST(0 AS DOUBLE)) WHEN (1 = 1) THEN CAST(1 AS DOUBLE) ELSE (CAST(2 AS DOUBLE) / CAST(0 AS DOUBLE)) END:double>
+struct<CASE WHEN (1 = 0) THEN (CAST(1 AS DOUBLE) / CAST(CAST(udf(0) AS DOUBLE) AS DOUBLE)) WHEN (1 = 1) THEN CAST(1 AS DOUBLE) ELSE (CAST(2 AS DOUBLE) / CAST(0 AS DOUBLE)) END:double>
-- !query 20 output
1.0
-- !query 21
-SELECT CASE WHEN i > 100 THEN 1/0 ELSE 0 END FROM case_tbl
+SELECT CASE WHEN i > 100 THEN udf(1/0) ELSE udf(0) END FROM case_tbl
-- !query 21 schema
-struct<CASE WHEN (i > 100) THEN (CAST(1 AS DOUBLE) / CAST(0 AS DOUBLE)) ELSE CAST(0 AS DOUBLE) END:double>
+struct<CASE WHEN (i > 100) THEN udf((cast(1 as double) / cast(0 as double))) ELSE udf(0) END:string>
-- !query 21 output
-0.0
-0.0
-0.0
-0.0
+0
+0
+0
+0
-- !query 22
-SELECT CASE 'a' WHEN 'a' THEN 1 ELSE 2 END
+SELECT CASE 'a' WHEN 'a' THEN udf(1) ELSE udf(2) END
-- !query 22 schema
-struct<CASE WHEN (a = a) THEN 1 ELSE 2 END:int>
+struct<CASE WHEN (a = a) THEN udf(1) ELSE udf(2) END:string>
-- !query 22 output
1
-283,7 +283,7 big
-- !query 27
-SELECT * FROM CASE_TBL WHERE COALESCE(f,i) = 4
+SELECT * FROM CASE_TBL WHERE udf(COALESCE(f,i)) = 4
-- !query 27 schema
struct<i:int,f:double>
-- !query 27 output
-291,7 +291,7 struct<i:int,f:double>
-- !query 28
-SELECT * FROM CASE_TBL WHERE NULLIF(f,i) = 2
+SELECT * FROM CASE_TBL WHERE udf(NULLIF(f,i)) = 2
-- !query 28 schema
struct<i:int,f:double>
-- !query 28 output
-299,10 +299,10 struct<i:int,f:double>
-- !query 29
-SELECT COALESCE(a.f, b.i, b.j)
+SELECT udf(COALESCE(a.f, b.i, b.j))
FROM CASE_TBL a, CASE2_TBL b
-- !query 29 schema
-struct<coalesce(f, CAST(i AS DOUBLE), CAST(j AS DOUBLE)):double>
+struct<udf(coalesce(f, cast(i as double), cast(j as double))):string>
-- !query 29 output
-30.3
-30.3
-332,8 +332,8 struct<coalesce(f, CAST(i AS DOUBLE), CAST(j AS DOUBLE)):double>
-- !query 30
SELECT *
- FROM CASE_TBL a, CASE2_TBL b
- WHERE COALESCE(a.f, b.i, b.j) = 2
+ FROM CASE_TBL a, CASE2_TBL b
+ WHERE udf(COALESCE(a.f, b.i, b.j)) = 2
-- !query 30 schema
struct<i:int,f:double,i:int,j:int>
-- !query 30 output
-342,7 +342,7 struct<i:int,f:double,i:int,j:int>
-- !query 31
-SELECT '' AS Five, NULLIF(a.i,b.i) AS `NULLIF(a.i,b.i)`,
+SELECT udf('') AS Five, NULLIF(a.i,b.i) AS `NULLIF(a.i,b.i)`,
NULLIF(b.i, 4) AS `NULLIF(b.i,4)`
FROM CASE_TBL a, CASE2_TBL b
-- !query 31 schema
-377,7 +377,7 struct<Five:string,NULLIF(a.i,b.i):int,NULLIF(b.i,4):int>
-- !query 32
SELECT '' AS `Two`, *
FROM CASE_TBL a, CASE2_TBL b
- WHERE COALESCE(f,b.i) = 2
+ WHERE CAST(udf(COALESCE(f,b.i) = 2) AS boolean)
-- !query 32 schema
struct<Two:string,i:int,f:double,i:int,j:int>
-- !query 32 output
-388,15 +388,15 struct<Two:string,i:int,f:double,i:int,j:int>
-- !query 33
SELECT CASE
(CASE vol('bar')
- WHEN 'foo' THEN 'it was foo!'
- WHEN vol(null) THEN 'null input'
+ WHEN udf('foo') THEN 'it was foo!'
+ WHEN udf(vol(null)) THEN 'null input'
WHEN 'bar' THEN 'it was bar!' END
)
- WHEN 'it was foo!' THEN 'foo recognized'
- WHEN 'it was bar!' THEN 'bar recognized'
- ELSE 'unrecognized' END
+ WHEN udf('it was foo!') THEN 'foo recognized'
+ WHEN 'it was bar!' THEN udf('bar recognized')
+ ELSE 'unrecognized' END AS col
-- !query 33 schema
-struct<CASE WHEN (CASE WHEN (UDF:vol(bar) = foo) THEN it was foo! WHEN (UDF:vol(bar) = UDF:vol(null)) THEN null input WHEN (UDF:vol(bar) = bar) THEN it was bar! END = it was foo!) THEN foo recognized WHEN (CASE WHEN (UDF:vol(bar) = foo) THEN it was foo! WHEN (UDF:vol(bar) = UDF:vol(null)) THEN null input WHEN (UDF:vol(bar) = bar) THEN it was bar! END = it was bar!) THEN bar recognized ELSE unrecognized END:string>
+struct<col:string>
-- !query 33 output
bar recognized
```
</p>
</details>
https://github.com/apache/spark/pull/25069 contains the same minor fixes as it's required to write the tests.
## How was this patch tested?
Tested as guided in [SPARK-27921](https://issues.apache.org/jira/browse/SPARK-27921).
Closes#25070 from HyukjinKwon/SPARK-28273.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: Wenchen Fan <wenchen@databricks.com>
## What changes were proposed in this pull request?
This PR proposes to rename `mapPartitionsInPandas` to `mapInPandas` with a separate evaluation type .
Had an offline discussion with rxin, mengxr and cloud-fan
The reason is basically:
1. `SCALAR_ITER` doesn't make sense with `mapPartitionsInPandas`.
2. It cannot share the same Pandas UDF, for instance, at `select` and `mapPartitionsInPandas` unlike `GROUPED_AGG` because iterator's return type is different.
3. `mapPartitionsInPandas` -> `mapInPandas` - see https://github.com/apache/spark/pull/25044#issuecomment-508298552 and https://github.com/apache/spark/pull/25044#issuecomment-508299764
Renaming `SCALAR_ITER` as `MAP_ITER` is abandoned due to 2. reason.
For `XXX_ITER`, it might have to have a different interface in the future if we happen to add other versions of them. But this is an orthogonal topic with `mapPartitionsInPandas`.
## How was this patch tested?
Existing tests should cover.
Closes#25044 from HyukjinKwon/SPARK-28198.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
In Python 2.7 with latest PyArrow and Pandas, the error message seems a bit different with Python 3. This PR simply fixes the test.
```
======================================================================
FAIL: test_createDataFrame_with_incorrect_schema (pyspark.sql.tests.test_arrow.ArrowTests)
----------------------------------------------------------------------
Traceback (most recent call last):
File "/.../spark/python/pyspark/sql/tests/test_arrow.py", line 275, in test_createDataFrame_with_incorrect_schema
self.spark.createDataFrame(pdf, schema=wrong_schema)
AssertionError: "integer.*required.*got.*str" does not match "('Exception thrown when converting pandas.Series (object) to Arrow Array (int32). It can be caused by overflows or other unsafe conversions warned by Arrow. Arrow safe type check can be disabled by using SQL config `spark.sql.execution.pandas.arrowSafeTypeConversion`.', ArrowTypeError('an integer is required',))"
======================================================================
FAIL: test_createDataFrame_with_incorrect_schema (pyspark.sql.tests.test_arrow.EncryptionArrowTests)
----------------------------------------------------------------------
Traceback (most recent call last):
File "/.../spark/python/pyspark/sql/tests/test_arrow.py", line 275, in test_createDataFrame_with_incorrect_schema
self.spark.createDataFrame(pdf, schema=wrong_schema)
AssertionError: "integer.*required.*got.*str" does not match "('Exception thrown when converting pandas.Series (object) to Arrow Array (int32). It can be caused by overflows or other unsafe conversions warned by Arrow. Arrow safe type check can be disabled by using SQL config `spark.sql.execution.pandas.arrowSafeTypeConversion`.', ArrowTypeError('an integer is required',))"
```
## How was this patch tested?
Manually tested.
```
cd python
./run-tests --python-executables=python --modules pyspark-sql
```
Closes#25042 from HyukjinKwon/SPARK-28240.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
This PR proposes to add `mapPartitionsInPandas` API to DataFrame by using existing `SCALAR_ITER` as below:
1. Filtering via setting the column
```python
from pyspark.sql.functions import pandas_udf, PandasUDFType
df = spark.createDataFrame([(1, 21), (2, 30)], ("id", "age"))
pandas_udf(df.schema, PandasUDFType.SCALAR_ITER)
def filter_func(iterator):
for pdf in iterator:
yield pdf[pdf.id == 1]
df.mapPartitionsInPandas(filter_func).show()
```
```
+---+---+
| id|age|
+---+---+
| 1| 21|
+---+---+
```
2. `DataFrame.loc`
```python
from pyspark.sql.functions import pandas_udf, PandasUDFType
import pandas as pd
df = spark.createDataFrame([['aa'], ['bb'], ['cc'], ['aa'], ['aa'], ['aa']], ["value"])
pandas_udf(df.schema, PandasUDFType.SCALAR_ITER)
def filter_func(iterator):
for pdf in iterator:
yield pdf.loc[pdf.value.str.contains('^a'), :]
df.mapPartitionsInPandas(filter_func).show()
```
```
+-----+
|value|
+-----+
| aa|
| aa|
| aa|
| aa|
+-----+
```
3. `pandas.melt`
```python
from pyspark.sql.functions import pandas_udf, PandasUDFType
import pandas as pd
df = spark.createDataFrame(
pd.DataFrame({'A': {0: 'a', 1: 'b', 2: 'c'},
'B': {0: 1, 1: 3, 2: 5},
'C': {0: 2, 1: 4, 2: 6}}))
pandas_udf("A string, variable string, value long", PandasUDFType.SCALAR_ITER)
def filter_func(iterator):
for pdf in iterator:
import pandas as pd
yield pd.melt(pdf, id_vars=['A'], value_vars=['B', 'C'])
df.mapPartitionsInPandas(filter_func).show()
```
```
+---+--------+-----+
| A|variable|value|
+---+--------+-----+
| a| B| 1|
| a| C| 2|
| b| B| 3|
| b| C| 4|
| c| B| 5|
| c| C| 6|
+---+--------+-----+
```
The current limitation of `SCALAR_ITER` is that it doesn't allow different length of result, which is pretty critical in practice - for instance, we cannot simply filter by using Pandas APIs but we merely just map N to N. This PR allows map N to M like flatMap.
This API mimics the way of `mapPartitions` but keeps API shape of `SCALAR_ITER` by allowing different results.
### How does this PR implement?
This PR adds mimics both `dapply` with Arrow optimization and Grouped Map Pandas UDF. At Python execution side, it reuses existing `SCALAR_ITER` code path.
Therefore, externally, we don't introduce any new type of Pandas UDF but internally we use another evaluation type code `205` (`SQL_MAP_PANDAS_ITER_UDF`).
This approach is similar with Pandas' Windows function implementation with Grouped Aggregation Pandas UDF functions - internally we have `203` (`SQL_WINDOW_AGG_PANDAS_UDF`) but externally we just share the same `GROUPED_AGG`.
## How was this patch tested?
Manually tested and unittests were added.
Closes#24997 from HyukjinKwon/scalar-udf-iter.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
The documentation in `linalg.py` is not consistent. This PR uniforms the documentation.
## How was this patch tested?
NA
Closes#25011 from mgaido91/SPARK-28170.
Authored-by: Marco Gaido <marcogaido91@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
Add docstring/doctest for `SCALAR_ITER` Pandas UDF. I explicitly mentioned that per-partition execution is an implementation detail, not guaranteed. I will submit another PR to add the same to user guide, just to keep this PR minimal.
I didn't add "doctest: +SKIP" in the first commit so it is easy to test locally.
cc: HyukjinKwon gatorsmile icexelloss BryanCutler WeichenXu123


## How was this patch tested?
doctest
Closes#25005 from mengxr/SPARK-28056.2.
Authored-by: Xiangrui Meng <meng@databricks.com>
Signed-off-by: gatorsmile <gatorsmile@gmail.com>
## What changes were proposed in this pull request?
Closes the generator when Python UDFs stop early.
### Manually verification on pandas iterator UDF and mapPartitions
```python
from pyspark.sql import SparkSession
from pyspark.sql.functions import pandas_udf, PandasUDFType
from pyspark.sql.functions import col, udf
from pyspark.taskcontext import TaskContext
import time
import os
spark.conf.set('spark.sql.execution.arrow.maxRecordsPerBatch', '1')
spark.conf.set('spark.sql.pandas.udf.buffer.size', '4')
pandas_udf("int", PandasUDFType.SCALAR_ITER)
def fi1(it):
try:
for batch in it:
yield batch + 100
time.sleep(1.0)
except BaseException as be:
print("Debug: exception raised: " + str(type(be)))
raise be
finally:
open("/tmp/000001.tmp", "a").close()
df1 = spark.range(10).select(col('id').alias('a')).repartition(1)
# will see log Debug: exception raised: <class 'GeneratorExit'>
# and file "/tmp/000001.tmp" generated.
df1.select(col('a'), fi1('a')).limit(2).collect()
def mapper(it):
try:
for batch in it:
yield batch
except BaseException as be:
print("Debug: exception raised: " + str(type(be)))
raise be
finally:
open("/tmp/000002.tmp", "a").close()
df2 = spark.range(10000000).repartition(1)
# will see log Debug: exception raised: <class 'GeneratorExit'>
# and file "/tmp/000002.tmp" generated.
df2.rdd.mapPartitions(mapper).take(2)
```
## How was this patch tested?
Unit test added.
Please review https://spark.apache.org/contributing.html before opening a pull request.
Closes#24986 from WeichenXu123/pandas_iter_udf_limit.
Authored-by: WeichenXu <weichen.xu@databricks.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
Currently with `toLocalIterator()` and `toPandas()` with Arrow enabled, if the Spark job being run in the background serving thread errors, it will be caught and sent to Python through the PySpark serializer.
This is not the ideal solution because it is only catch a SparkException, it won't handle an error that occurs in the serializer, and each method has to have it's own special handling to propagate the error.
This PR instead returns the Python Server object along with the serving port and authentication info, so that it allows the Python caller to join with the serving thread. During the call to join, the serving thread Future is completed either successfully or with an exception. In the latter case, the exception will be propagated to Python through the Py4j call.
## How was this patch tested?
Existing tests
Closes#24834 from BryanCutler/pyspark-propagate-server-error-SPARK-27992.
Authored-by: Bryan Cutler <cutlerb@gmail.com>
Signed-off-by: Bryan Cutler <cutlerb@gmail.com>
## What changes were proposed in this pull request?
add missing RankingEvaluator
## How was this patch tested?
added testsuites
Closes#24869 from zhengruifeng/ranking_eval.
Authored-by: zhengruifeng <ruifengz@foxmail.com>
Signed-off-by: Sean Owen <sean.owen@databricks.com>
## What changes were proposed in this pull request?
This patch removes `fillna(0)` when creating ArrowBatch from a pandas Series.
With `fillna(0)`, the original code would turn a timestamp type into object type, which pyarrow will complain later:
```
>>> s = pd.Series([pd.NaT, pd.Timestamp('2015-01-01')])
>>> s.dtypes
dtype('<M8[ns]')
>>> s.fillna(0)
0 0
1 2015-01-01 00:00:00
dtype: object
```
## How was this patch tested?
Added `test_timestamp_nat`
Closes#24844 from icexelloss/SPARK-28003-arrow-nat.
Authored-by: Li Jin <ice.xelloss@gmail.com>
Signed-off-by: Bryan Cutler <cutlerb@gmail.com>
## What changes were proposed in this pull request?
Currently, pretty skipped message added by f7435bec6a mechanism seems not working when xmlrunner is installed apparently.
This PR fixes two things:
1. When `xmlrunner` is installed, seems `xmlrunner` does not respect `vervosity` level in unittests (default is level 1).
So the output looks as below
```
Running tests...
----------------------------------------------------------------------
SSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSS
----------------------------------------------------------------------
```
So it is not caught by our message detection mechanism.
2. If we manually set the `vervocity` level to `xmlrunner`, it prints messages as below:
```
test_mixed_udf (pyspark.sql.tests.test_pandas_udf_scalar.ScalarPandasUDFTests) ... SKIP (0.000s)
test_mixed_udf_and_sql (pyspark.sql.tests.test_pandas_udf_scalar.ScalarPandasUDFTests) ... SKIP (0.000s)
...
```
This is different in our Jenkins machine:
```
test_createDataFrame_column_name_encoding (pyspark.sql.tests.test_arrow.ArrowTests) ... skipped 'Pandas >= 0.23.2 must be installed; however, it was not found.'
test_createDataFrame_does_not_modify_input (pyspark.sql.tests.test_arrow.ArrowTests) ... skipped 'Pandas >= 0.23.2 must be installed; however, it was not found.'
...
```
Note that last `SKIP` is different. This PR fixes the regular expression to catch `SKIP` case as well.
## How was this patch tested?
Manually tested.
**Before:**
```
Starting test(python2.7): pyspark....
Finished test(python2.7): pyspark.... (0s)
...
Tests passed in 562 seconds
========================================================================
...
```
**After:**
```
Starting test(python2.7): pyspark....
Finished test(python2.7): pyspark.... (48s) ... 93 tests were skipped
...
Tests passed in 560 seconds
Skipped tests pyspark.... with python2.7:
pyspark...(...) ... SKIP (0.000s)
...
========================================================================
...
```
Closes#24927 from HyukjinKwon/SPARK-28130.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
When running FlatMapGroupsInPandasExec or AggregateInPandasExec the shuffle uses a default number of partitions of 200 in "spark.sql.shuffle.partitions". If the data is small, e.g. in testing, many of the partitions will be empty but are treated just the same.
This PR checks the `mapPartitionsInternal` iterator to be non-empty before calling `ArrowPythonRunner` to start computation on the iterator.
## How was this patch tested?
Existing tests. Ran the following benchmarks a simple example where most partitions are empty:
```python
from pyspark.sql.functions import pandas_udf, PandasUDFType
from pyspark.sql.types import *
df = spark.createDataFrame(
[(1, 1.0), (1, 2.0), (2, 3.0), (2, 5.0), (2, 10.0)],
("id", "v"))
pandas_udf("id long, v double", PandasUDFType.GROUPED_MAP)
def normalize(pdf):
v = pdf.v
return pdf.assign(v=(v - v.mean()) / v.std())
df.groupby("id").apply(normalize).count()
```
**Before**
```
In [4]: %timeit df.groupby("id").apply(normalize).count()
1.58 s ± 62.8 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
In [5]: %timeit df.groupby("id").apply(normalize).count()
1.52 s ± 29.5 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
In [6]: %timeit df.groupby("id").apply(normalize).count()
1.52 s ± 37.8 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
```
**After this Change**
```
In [2]: %timeit df.groupby("id").apply(normalize).count()
646 ms ± 89.9 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
In [3]: %timeit df.groupby("id").apply(normalize).count()
408 ms ± 84.6 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
In [4]: %timeit df.groupby("id").apply(normalize).count()
381 ms ± 29.9 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
```
Closes#24926 from BryanCutler/pyspark-pandas_udf-map-agg-skip-empty-parts-SPARK-28128.
Authored-by: Bryan Cutler <cutlerb@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
expose more metrics in evaluator: weightedTruePositiveRate/weightedFalsePositiveRate/weightedFMeasure/truePositiveRateByLabel/falsePositiveRateByLabel/precisionByLabel/recallByLabel/fMeasureByLabel
## How was this patch tested?
existing cases and add cases
Closes#24868 from zhengruifeng/multi_class_support_bylabel.
Authored-by: zhengruifeng <ruifengz@foxmail.com>
Signed-off-by: Sean Owen <sean.owen@databricks.com>
## What changes were proposed in this pull request?
When using `DROPMALFORMED` mode, corrupted records aren't dropped if malformed columns aren't read. This behavior is due to CSV parser column pruning. Current doc of `DROPMALFORMED` doesn't mention the effect of column pruning. Users will be confused by the fact that `DROPMALFORMED` mode doesn't work as expected.
Column pruning also affects other modes. This is a doc improvement to add a note to doc of `mode` to explain it.
## How was this patch tested?
N/A. This is just doc change.
Closes#24894 from viirya/SPARK-28058.
Authored-by: Liang-Chi Hsieh <viirya@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
This increases the minimum supported version of Pandas to 0.23.2. Using a lower version will raise an error `Pandas >= 0.23.2 must be installed; however, your version was 0.XX`. Also, a workaround for using pyarrow with Pandas 0.19.2 was removed.
## How was this patch tested?
Existing Tests
Closes#24867 from BryanCutler/pyspark-increase-min-pandas-SPARK-28041.
Authored-by: Bryan Cutler <cutlerb@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
Allow Pandas UDF to take an iterator of pd.Series or an iterator of tuple of pd.Series.
Note the UDF input args will be always one iterator:
* if the udf take only column as input, the iterator's element will be pd.Series (corresponding to the column values batch)
* if the udf take multiple columns as inputs, the iterator's element will be a tuple composed of multiple `pd.Series`s, each one corresponding to the multiple columns as inputs (keep the same order). For example:
```
pandas_udf("int", PandasUDFType.SCALAR_ITER)
def the_udf(iterator):
for col1_batch, col2_batch in iterator:
yield col1_batch + col2_batch
df.select(the_udf("col1", "col2"))
```
The udf above will add col1 and col2.
I haven't add unit tests, but manually tests show it works fine. So it is ready for first pass review.
We can test several typical cases:
```
from pyspark.sql import SparkSession
from pyspark.sql.functions import pandas_udf, PandasUDFType
from pyspark.sql.functions import udf
from pyspark.taskcontext import TaskContext
df = spark.createDataFrame([(1, 20), (3, 40)], ["a", "b"])
pandas_udf("int", PandasUDFType.SCALAR_ITER)
def fi1(it):
pid = TaskContext.get().partitionId()
print("DBG: fi1: do init stuff, partitionId=" + str(pid))
for batch in it:
yield batch + 100
print("DBG: fi1: do close stuff, partitionId=" + str(pid))
pandas_udf("int", PandasUDFType.SCALAR_ITER)
def fi2(it):
pid = TaskContext.get().partitionId()
print("DBG: fi2: do init stuff, partitionId=" + str(pid))
for batch in it:
yield batch + 10000
print("DBG: fi2: do close stuff, partitionId=" + str(pid))
pandas_udf("int", PandasUDFType.SCALAR_ITER)
def fi3(it):
pid = TaskContext.get().partitionId()
print("DBG: fi3: do init stuff, partitionId=" + str(pid))
for x, y in it:
yield x + y * 10 + 100000
print("DBG: fi3: do close stuff, partitionId=" + str(pid))
pandas_udf("int", PandasUDFType.SCALAR)
def fp1(x):
return x + 1000
udf("int")
def fu1(x):
return x + 10
# test select "pandas iter udf/pandas udf/sql udf" expressions at the same time.
# Note this case the `fi1("a"), fi2("b"), fi3("a", "b")` will generate only one plan,
# and `fu1("a")`, `fp1("a")` will generate another two separate plans.
df.select(fi1("a"), fi2("b"), fi3("a", "b"), fu1("a"), fp1("a")).show()
# test chain two pandas iter udf together
# Note this case `fi2(fi1("a"))` will generate only one plan
# Also note the init stuff/close stuff call order will be like:
# (debug output following)
# DBG: fi2: do init stuff, partitionId=0
# DBG: fi1: do init stuff, partitionId=0
# DBG: fi1: do close stuff, partitionId=0
# DBG: fi2: do close stuff, partitionId=0
df.select(fi2(fi1("a"))).show()
# test more complex chain
# Note this case `fi1("a"), fi2("a")` will generate one plan,
# and `fi3(fi1_output, fi2_output)` will generate another plan
df.select(fi3(fi1("a"), fi2("a"))).show()
```
## How was this patch tested?
To be added.
Please review http://spark.apache.org/contributing.html before opening a pull request.
Closes#24643 from WeichenXu123/pandas_udf_iter.
Lead-authored-by: WeichenXu <weichen.xu@databricks.com>
Co-authored-by: Xiangrui Meng <meng@databricks.com>
Signed-off-by: Xiangrui Meng <meng@databricks.com>
## What changes were proposed in this pull request?
I suspect that the doc of `rowsBetween` methods in Scala and PySpark looks wrong.
Because:
```scala
scala> val df = Seq((1, "a"), (2, "a"), (3, "a"), (4, "a"), (5, "a"), (6, "a")).toDF("id", "category")
df: org.apache.spark.sql.DataFrame = [id: int, category: string]
scala> val byCategoryOrderedById = Window.partitionBy('category).orderBy('id).rowsBetween(-1, 2)
byCategoryOrderedById: org.apache.spark.sql.expressions.WindowSpec = org.apache.spark.sql.expressions.WindowSpec7f04de97
scala> df.withColumn("sum", sum('id) over byCategoryOrderedById).show()
+---+--------+---+
| id|category|sum|
+---+--------+---+
| 1| a| 6| # sum from index 0 to (0 + 2): 1 + 2 + 3 = 6
| 2| a| 10| # sum from index (1 - 1) to (1 + 2): 1 + 2 + 3 + 4 = 10
| 3| a| 14|
| 4| a| 18|
| 5| a| 15|
| 6| a| 11|
+---+--------+---+
```
So the frame (-1, 2) for row with index 5, as described in the doc, should range from index 4 to index 7.
## How was this patch tested?
N/A, just doc change.
Closes#24864 from viirya/window-spec-doc.
Authored-by: Liang-Chi Hsieh <viirya@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
add MultilabelClassificationEvaluator
## How was this patch tested?
added testsuites
Closes#24777 from zhengruifeng/multi_label_eval.
Authored-by: zhengruifeng <ruifengz@foxmail.com>
Signed-off-by: Sean Owen <sean.owen@databricks.com>
## What changes were proposed in this pull request?
Just found the doctest on `over` function of `Column` is commented out. The window spec is also not for the window function used there.
We should either remove the doctest, or improve it.
Because other functions of `Column` have doctest generally, so this PR tries to improve it.
## How was this patch tested?
Added doctest.
Closes#24854 from viirya/column-test-minor.
Authored-by: Liang-Chi Hsieh <viirya@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
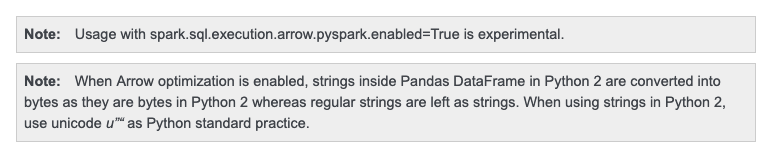
When Arrow optimization is enabled in Python 2.7,
```python
import pandas
pdf = pandas.DataFrame(["test1", "test2"])
df = spark.createDataFrame(pdf)
df.show()
```
I got the following output:
```
+----------------+
| 0|
+----------------+
|[74 65 73 74 31]|
|[74 65 73 74 32]|
+----------------+
```
This looks because Python's `str` and `byte` are same. it does look right:
```python
>>> str == bytes
True
>>> isinstance("a", bytes)
True
```
To cut it short:
1. Python 2 treats `str` as `bytes`.
2. PySpark added some special codes and hacks to recognizes `str` as string types.
3. PyArrow / Pandas followed Python 2 difference
To fix, we have two options:
1. Fix it to match the behaviour to PySpark's
2. Note the differences
but Python 2 is deprecated anyway. I think it's better to just note it and for go option 2.
## How was this patch tested?
Manually tested.
Doc was checked too:
Closes#24838 from HyukjinKwon/SPARK-27995.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
Flush batch timely for pandas UDF.
This could improve performance when multiple pandas UDF plans are pipelined.
When batch being flushed in time, downstream pandas UDFs will get pipelined as soon as possible, and pipeline will help hide the donwstream UDFs computation time. For example:
When the first UDF start computing on batch-3, the second pipelined UDF can start computing on batch-2, and the third pipelined UDF can start computing on batch-1.
If we do not flush each batch in time, the donwstream UDF's pipeline will lag behind too much, which may increase the total processing time.
I add flush at two places:
* JVM process feed data into python worker. In jvm side, when write one batch, flush it
* VM process read data from python worker output, In python worker side, when write one batch, flush it
If no flush, the default buffer size for them are both 65536. Especially in the ML case, in order to make realtime prediction, we will make batch size very small. The buffer size is too large for the case, which cause downstream pandas UDF pipeline lag behind too much.
### Note
* This is only applied to pandas scalar UDF.
* Do not flush for each batch. The minimum interval between two flush is 0.1 second. This avoid too frequent flushing when batch size is small. It works like:
```
last_flush_time = time.time()
for batch in iterator:
writer.write_batch(batch)
flush_time = time.time()
if self.flush_timely and (flush_time - last_flush_time > 0.1):
stream.flush()
last_flush_time = flush_time
```
## How was this patch tested?
### Benchmark to make sure the flush do not cause performance regression
#### Test code:
```
numRows = ...
batchSize = ...
spark.conf.set('spark.sql.execution.arrow.maxRecordsPerBatch', str(batchSize))
df = spark.range(1, numRows + 1, numPartitions=1).select(col('id').alias('a'))
pandas_udf("int", PandasUDFType.SCALAR)
def fp1(x):
return x + 10
beg_time = time.time()
result = df.select(sum(fp1('a'))).head()
print("result: " + str(result[0]))
print("consume time: " + str(time.time() - beg_time))
```
#### Test Result:
params | Consume time (Before) | Consume time (After)
------------ | ----------------------- | ----------------------
numRows=100000000, batchSize=10000 | 23.43s | 24.64s
numRows=100000000, batchSize=1000 | 36.73s | 34.50s
numRows=10000000, batchSize=100 | 35.67s | 32.64s
numRows=1000000, batchSize=10 | 33.60s | 32.11s
numRows=100000, batchSize=1 | 33.36s | 31.82s
### Benchmark pipelined pandas UDF
#### Test code:
```
spark.conf.set('spark.sql.execution.arrow.maxRecordsPerBatch', '1')
df = spark.range(1, 31, numPartitions=1).select(col('id').alias('a'))
pandas_udf("int", PandasUDFType.SCALAR)
def fp1(x):
print("run fp1")
time.sleep(1)
return x + 100
pandas_udf("int", PandasUDFType.SCALAR)
def fp2(x, y):
print("run fp2")
time.sleep(1)
return x + y
beg_time = time.time()
result = df.select(sum(fp2(fp1('a'), col('a')))).head()
print("result: " + str(result[0]))
print("consume time: " + str(time.time() - beg_time))
```
#### Test Result:
**Before**: consume time: 63.57s
**After**: consume time: 32.43s
**So the PR improve performance by make downstream UDF get pipelined early.**
Please review https://spark.apache.org/contributing.html before opening a pull request.
Closes#24734 from WeichenXu123/improve_pandas_udf_pipeline.
Lead-authored-by: WeichenXu <weichen.xu@databricks.com>
Co-authored-by: Xiangrui Meng <meng@databricks.com>
Signed-off-by: gatorsmile <gatorsmile@gmail.com>
## What changes were proposed in this pull request?
Similar to https://github.com/apache/spark/pull/24070, we now propagate SparkExceptions that are encountered during the collect in the java process to the python process.
Fixes https://jira.apache.org/jira/browse/SPARK-27805
## How was this patch tested?
Added a new unit test
Closes#24677 from dvogelbacher/dv/betterErrorMsgWhenUsingArrow.
Authored-by: David Vogelbacher <dvogelbacher@palantir.com>
Signed-off-by: Bryan Cutler <cutlerb@gmail.com>
## What changes were proposed in this pull request?
Added support for `*` and `^` operators, along with expressions within parentheses. New operators just expand to already supported terms, such as;
- y ~ a * b = y ~ a + b + a : b
- y ~ (a+b+c)^3 = y ~ a + b + c + a : b + a : c + a :b : c
## How was this patch tested?
Added new unit tests to RFormulaParserSuite
mengxr yanboliang
Closes#24764 from ozancicek/rformula.
Authored-by: ozan <ozancancicekci@gmail.com>
Signed-off-by: Sean Owen <sean.owen@databricks.com>
## What changes were proposed in this pull request?
Add deprecation warning for Python 2.
## How was this patch tested?
Manual tests:
Interactive shell:
~~~
$ bin/pyspark
Python 2.7.15 |Anaconda, Inc.| (default, Nov 13 2018, 17:07:45)
[GCC 4.2.1 Compatible Clang 4.0.1 (tags/RELEASE_401/final)] on darwin
Type "help", "copyright", "credits" or "license" for more information.
19/06/03 14:54:13 WARN NativeCodeLoader: Unable to load native-hadoop library for your platform... using builtin-java classes where applicable
/Users/meng/src/spark/python/pyspark/context.py:219: DeprecationWarning: Support for Python 2 is deprecated as of Spark 3.0. See the plan for dropping Python 2 support at https://spark.apache.org/news/plan-for-dropping-python-2-support.html.
DeprecationWarning)
Welcome to
____ __
/ __/__ ___ _____/ /__
_\ \/ _ \/ _ `/ __/ '_/
/__ / .__/\_,_/_/ /_/\_\ version 3.0.0-SNAPSHOT
/_/
Using Python version 2.7.15 (default, Nov 13 2018 17:07:45)
SparkSession available as 'spark'.
>>>
~~~
spark-submit job (with default log level set to WARN):
~~~
$ bin/spark-submit test.py
19/06/03 14:54:38 WARN NativeCodeLoader: Unable to load native-hadoop library for your platform... using builtin-java classes where applicable
/Users/meng/src/spark/python/lib/pyspark.zip/pyspark/context.py:219: DeprecationWarning: Support for Python 2 is deprecated as of Spark 3.0. See the plan for dropping Python 2 support at https://spark.apache.org/news/plan-for-dropping-python-2-support.html.
DeprecationWarning)
~~~
Verified that warning messages do not show up in Python 3.
` DeprecationWarning)` is displayed at the end because `warn` by default print the code line. This behavior can be changed by monkey patching `showwarning` https://stackoverflow.com/questions/2187269/print-only-the-message-on-warnings. It might not worth the effort.
Closes#24786 from mengxr/SPARK-27887.
Authored-by: Xiangrui Meng <meng@databricks.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
`spark.sql.execution.arrow.enabled` was added when we add PySpark arrow optimization.
Later, in the current master, SparkR arrow optimization was added and it's controlled by the same configuration `spark.sql.execution.arrow.enabled`.
There look two issues about this:
1. `spark.sql.execution.arrow.enabled` in PySpark was added from 2.3.0 whereas SparkR optimization was added 3.0.0. The stability is different so it's problematic when we change the default value for one of both optimization first.
2. Suppose users want to share some JVM by PySpark and SparkR. They are currently forced to use the optimization for all or none if the configuration is set globally.
This PR proposes two separate configuration groups for PySpark and SparkR about Arrow optimization:
- Deprecate `spark.sql.execution.arrow.enabled`
- Add `spark.sql.execution.arrow.pyspark.enabled` (fallback to `spark.sql.execution.arrow.enabled`)
- Add `spark.sql.execution.arrow.sparkr.enabled`
- Deprecate `spark.sql.execution.arrow.fallback.enabled`
- Add `spark.sql.execution.arrow.pyspark.fallback.enabled ` (fallback to `spark.sql.execution.arrow.fallback.enabled`)
Note that `spark.sql.execution.arrow.maxRecordsPerBatch` is used within JVM side for both.
Note that `spark.sql.execution.arrow.fallback.enabled` was added due to behaviour change. We don't need it in SparkR - SparkR side has the automatic fallback.
## How was this patch tested?
Manually tested and some unittests were added.
Closes#24700 from HyukjinKwon/separate-sparkr-arrow.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
Unset InputFileBlockHolder at the end of tasks to stop the file name from leaking over to other tasks in the same thread. This happens in particular in Pyspark because of its complex threading model.
## How was this patch tested?
new pyspark test
Closes#24605 from jose-torres/fix254.
Authored-by: Jose Torres <torres.joseph.f+github@gmail.com>
Signed-off-by: Xingbo Jiang <xingbo.jiang@databricks.com>
## What changes were proposed in this pull request?
https://github.com/apache/spark/pull/22275 introduced a performance improvement where we send partitions out of order to python and then, as a last step, send the partition order as well.
However, if there are no partitions we will never send the partition order and we will get an "EofError" on the python side.
This PR fixes this by also sending the partition order if there are no partitions present.
## How was this patch tested?
New unit test added.
Closes#24650 from dvogelbacher/dv/fixNoPartitionArrowConversion.
Authored-by: David Vogelbacher <dvogelbacher@palantir.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
Tighten up some key links to the project and download pages to use HTTPS
## How was this patch tested?
N/A
Closes#24665 from srowen/HTTPSURLs.
Authored-by: Sean Owen <sean.owen@databricks.com>
Signed-off-by: Dongjoon Hyun <dhyun@apple.com>
## What changes were proposed in this pull request?
Not a big deal but it bugged me. This PR removes printing out plans in PySpark UDF tests.
Before:
```
Running tests...
----------------------------------------------------------------------
Setting default log level to "WARN".
To adjust logging level use sc.setLogLevel(newLevel). For SparkR, use setLogLevel(newLevel).
== Parsed Logical Plan ==
GlobalLimit 1
+- LocalLimit 1
+- Project [id#668L, <lambda>(id#668L) AS copy#673]
+- Sort [id#668L ASC NULLS FIRST], true
+- Range (0, 10, step=1, splits=Some(4))
== Analyzed Logical Plan ==
id: bigint, copy: int
GlobalLimit 1
+- LocalLimit 1
+- Project [id#668L, <lambda>(id#668L) AS copy#673]
+- Sort [id#668L ASC NULLS FIRST], true
+- Range (0, 10, step=1, splits=Some(4))
== Optimized Logical Plan ==
GlobalLimit 1
+- LocalLimit 1
+- Project [id#668L, pythonUDF0#676 AS copy#673]
+- BatchEvalPython [<lambda>(id#668L)], [id#668L, pythonUDF0#676]
+- Range (0, 10, step=1, splits=Some(4))
== Physical Plan ==
CollectLimit 1
+- *(2) Project [id#668L, pythonUDF0#676 AS copy#673]
+- BatchEvalPython [<lambda>(id#668L)], [id#668L, pythonUDF0#676]
+- *(1) Range (0, 10, step=1, splits=4)
...........................................
----------------------------------------------------------------------
Ran 43 tests in 19.777s
```
After:
```
Running tests...
----------------------------------------------------------------------
Setting default log level to "WARN".
To adjust logging level use sc.setLogLevel(newLevel). For SparkR, use setLogLevel(newLevel).
...........................................
----------------------------------------------------------------------
Ran 43 tests in 25.201s
```
## How was this patch tested?
N/A
Closes#24661 from HyukjinKwon/remove-explain-in-test.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
We updated our website a long time ago to describe Spark as the unified analytics engine, which is also how Spark is described in the community now. But our README and docs page still use the same description from 2011 ... This patch updates them.
The patch also updates the README example to use more modern APIs, and refer to Structured Streaming rather than Spark Streaming.
Closes#24573 from rxin/consistent-message.
Authored-by: Reynold Xin <rxin@databricks.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
Added method 'meanAveragePrecisionAt' k to RankingMetrics.
This branch is rebased with squashed commits from https://github.com/apache/spark/pull/24458
## How was this patch tested?
Added code in the existing test RankingMetricsSuite.
Please review http://spark.apache.org/contributing.html before opening a pull request.
Closes#24543 from qb-tarushg/SPARK-27540-REBASE.
Authored-by: qb-tarushg <tarush.grover@quantumblack.com>
Signed-off-by: Sean Owen <sean.owen@databricks.com>
## What changes were proposed in this pull request?
### Background:
The data source option `pathGlobFilter` is introduced for Binary file format: https://github.com/apache/spark/pull/24354 , which can be used for filtering file names, e.g. reading `.png` files only while there is `.json` files in the same directory.
### Proposal:
Make the option `pathGlobFilter` as a general option for all file sources. The path filtering should happen in the path globbing on Driver.
### Motivation:
Filtering the file path names in file scan tasks on executors is kind of ugly.
### Impact:
1. The splitting of file partitions will be more balanced.
2. The metrics of file scan will be more accurate.
3. Users can use the option for reading other file sources.
## How was this patch tested?
Unit tests
Closes#24518 from gengliangwang/globFilter.
Authored-by: Gengliang Wang <gengliang.wang@databricks.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
This fixes an error when a PySpark local iterator, for both RDD and DataFrames, goes out of scope and the connection is closed before fully consuming the iterator. The error occurs on the JVM in the serving thread, when Python closes the local socket while the JVM is writing to it. This usually happens when there is enough data to fill the socket read buffer, causing the write call to block.
Additionally, this fixes a problem when an error occurs in the Python worker and the collect job is cancelled with an exception. Previously, the Python driver was never notified of the error so the user could get a partial result (iteration until the error) and the application will continue. With this change, an error in the worker is sent to the Python iterator and is then raised.
The change here introduces a protocol for PySpark local iterators that work as follows:
1) The local socket connection is made when the iterator is created
2) When iterating, Python first sends a request for partition data as a non-zero integer
3) While the JVM local iterator over partitions has next, it triggers a job to collect the next partition
4) The JVM sends a nonzero response to indicate it has the next partition to send
5) The next partition is sent to Python and read by the PySpark deserializer
6) After sending the entire partition, an `END_OF_DATA_SECTION` is sent to Python which stops the deserializer and allows to make another request
7) When the JVM gets a request from Python but has already consumed it's local iterator, it will send a zero response to Python and both will close the socket cleanly
8) If an error occurs in the worker, a negative response is sent to Python followed by the error message. Python will then raise a RuntimeError with the message, stopping iteration.
9) When the PySpark local iterator is garbage-collected, it will read any remaining data from the current partition (this is data that has already been collected) and send a request of zero to tell the JVM to stop collection jobs and close the connection.
Steps 1, 3, 5, 6 are the same as before. Step 8 was completely missing before because errors in the worker were never communicated back to Python. The other steps add synchronization to allow for a clean closing of the socket, with a small trade-off in performance for each partition. This is mainly because the JVM does not start collecting partition data until it receives a request to do so, where before it would eagerly write all data until the socket receive buffer is full.
## How was this patch tested?
Added new unit tests for DataFrame and RDD `toLocalIterator` and tested not fully consuming the iterator. Manual tests with Python 2.7 and 3.6.
Closes#24070 from BryanCutler/pyspark-toLocalIterator-clean-stop-SPARK-23961.
Authored-by: Bryan Cutler <cutlerb@gmail.com>
Signed-off-by: Bryan Cutler <cutlerb@gmail.com>
This is PR is meant to replace #20503, which lay dormant for a while. The solution in the original PR is still valid, so this is just that patch rebased onto the current master.
Original summary follows.
## What changes were proposed in this pull request?
Fix `__repr__` behaviour for Rows.
Rows `__repr__` assumes data is a string when column name is missing.
Examples,
```
>>> from pyspark.sql.types import Row
>>> Row ("Alice", "11")
<Row(Alice, 11)>
>>> Row (name="Alice", age=11)
Row(age=11, name='Alice')
>>> Row ("Alice", 11)
<snip stack trace>
TypeError: sequence item 1: expected string, int found
```
This is because Row () when called without column names assumes everything is a string.
## How was this patch tested?
Manually tested and a unit test was added to `python/pyspark/sql/tests/test_types.py`.
Closes#24448 from tbcs/SPARK-23299.
Lead-authored-by: Tibor Csögör <tibi@tiborius.net>
Co-authored-by: Shashwat Anand <me@shashwat.me>
Signed-off-by: Holden Karau <holden@pigscanfly.ca>
## What changes were proposed in this pull request?
In SPARK-27612, one correctness issue was reported. When protocol 4 is used to pickle Python objects, we found that unpickled objects were wrong. A temporary fix was proposed by not using highest protocol.
It was found that Opcodes.MEMOIZE was appeared in the opcodes in protocol 4. It is suspect to this issue.
A deeper dive found that Opcodes.MEMOIZE stores objects into internal map of Unpickler object. We use single Unpickler object to unpickle serialized Python bytes. Stored objects intervenes next round of unpickling, if the map is not cleared.
We has two options:
1. Continues to reuse Unpickler, but calls its close after each unpickling.
2. Not to reuse Unpickler and create new Unpickler object in each unpickling.
This patch takes option 1.
## How was this patch tested?
Passing the test added in SPARK-27612 (#24519).
Closes#24521 from viirya/SPARK-27629.
Authored-by: Liang-Chi Hsieh <viirya@gmail.com>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
This PR partially reverts https://github.com/apache/spark/pull/20691
After we changed the Python protocol to highest ones, seems like it introduced a correctness bug. This potentially affects all Python related code paths.
I suspect a bug related to Pryolite (maybe opcodes `MEMOIZE`, `FRAME` and/or our `RowPickler`). I would like to stick to default protocol for now and investigate the issue separately.
I will separately investigate later to bring highest protocol back.
## How was this patch tested?
Unittest was added.
```bash
./run-tests --python-executables=python3.7 --testname "pyspark.sql.tests.test_serde SerdeTests.test_int_array_serialization"
```
Closes#24519 from HyukjinKwon/SPARK-27612.
Authored-by: HyukjinKwon <gurwls223@apache.org>
Signed-off-by: HyukjinKwon <gurwls223@apache.org>
## What changes were proposed in this pull request?
Add a non-intrusive button for python API documentation, which will remove ">>>" prompts and outputs of code - for easier copying of code.
For example: The below code-snippet in the document is difficult to copy due to ">>>" prompts
```
>>> l = [('Alice', 1)]
>>> spark.createDataFrame(l).collect()
[Row(_1='Alice', _2=1)]
```
Becomes this - After the copybutton in the corner of of code-block is pressed - which is easier to copy
```
l = [('Alice', 1)]
spark.createDataFrame(l).collect()
```
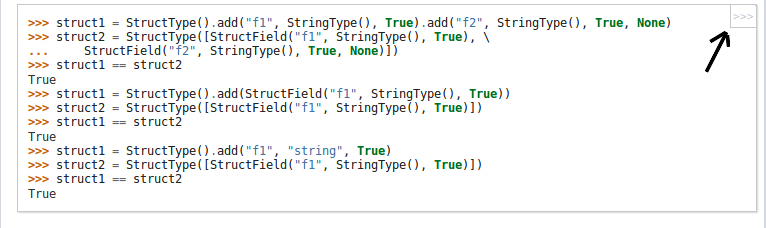
## File changes
Made changes to python/docs/conf.py and copybutton.js - thus only modifying sphinx frontend and no changes were made to the documentation itself- Build process for documentation remains the same.
copybutton.js -> This JS snippet was taken from the official python.org documentation site.
## How was this patch tested?
NA
Closes#24456 from sangramga/copybutton.
Authored-by: sangramga <sangramga@gmail.com>
Signed-off-by: Sean Owen <sean.owen@databricks.com>